Abstract
Background
Multiple common variants identified by genome-wide association studies showed limited evidence of the risk of breast cancer in Taiwan. In this study, we analyzed the breast cancer risk in relation to 13 individual single-nucleotide polymorphisms (SNPs) identified by a GWAS in an Asian population.
Methods
In total, 446 breast cancer patients and 514 healthy controls were recruited for this case–control study. In addition, we developed a polygenic risk score (PRS) including those variants significantly associated with breast cancer risk, and also evaluated the contribution of PRS and clinical risk factors to breast cancer using receiver operating characteristic curve (AUC).
Results
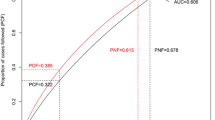
Logistic regression results showed that nine individual SNPs were significantly associated with breast cancer risk after multiple testing. Among all SNPs, six variants, namely FGFR2 (rs2981582), HCN1 (rs981782), MAP3K1 (rs889312), TOX3 (rs3803662), ZNF365 (rs10822013), and RAD51B (rs3784099), were selected to create PRS model. A dose–response association was observed between breast cancer risk and the PRS. Women in the highest quartile of PRS had a significantly increased risk compared to women in the lowest quartile (odds ratio 2.26; 95% confidence interval 1.51–3.38). The AUC for a model which contained the PRS in addition to clinical risk factors was 66.52%, whereas that for a model which with established risk factors only was 63.38%.
Conclusions
Our data identified a genetic risk predictor of breast cancer in Taiwanese population and suggest that risk models including PRS and clinical risk factors are useful in discriminating women at high risk of breast cancer from those at low risk.


Similar content being viewed by others
References
Health Promotion Administration MOHAW. Health indicator 123. https://olap.hpa.gov.tw/en_US/index.aspx. Accessed May 5 2016
Moller S, Mucci LA, Harris JR et al (2016) The heritability of breast cancer among women in the nordic twin study of cancer. Cancer Epidemiol Biomark Prev 25:145–150
Cai Q, Long J, Lu W et al (2011) Genome-wide association study identifies breast cancer risk variant at 10q21.2: results from the Asia Breast Cancer Consortium. Hum Mol Genet 20:4991–4999
Easton DF, Pooley KA, Dunning AM et al (2007) Genome-wide association study identifies novel breast cancer susceptibility loci. Nature 447:1087–1093
Gold B, Kirchhoff T, Stefanov S et al (2008) Genome-wide association study provides evidence for a breast cancer risk locus at 6q22.33. Proc Natl Acad Sci USA 105:4340–4345
Hunter DJ, Kraft P, Jacobs KB et al (2007) A genome-wide association study identifies alleles in FGFR2 associated with risk of sporadic postmenopausal breast cancer. Nat Genet 39:870–874
Kim HC, Lee JY, Sung H et al (2012) A genome-wide association study identifies a breast cancer risk variant in ERBB4 at 2q34: results from the Seoul Breast Cancer Study. Breast Cancer Res 14:R56
Long J, Cai Q, Sung H et al (2012) Genome-wide association study in East Asians identifies novel susceptibility loci for breast cancer. PLoS Genet 8:e1002532
Stacey SN, Manolescu A, Sulem P et al (2007) Common variants on chromosomes 2q35 and 16q12 confer susceptibility to estrogen receptor-positive breast cancer. Nat Genet 39:865–869
Stacey SN, Manolescu A, Sulem P et al (2008) Common variants on chromosome 5p12 confer susceptibility to estrogen receptor-positive breast cancer. Nat Genet 40:703–706
Mavaddat N, Antoniou AC, Easton DF, Garcia-Closas M (2010) Genetic susceptibility to breast cancer. Mol Oncol 4:174–191
Peto J, Collins N, Barfoot R et al (1999) Prevalence of BRCA1 and BRCA2 gene mutations in patients with early-onset breast cancer. J Natl Cancer Inst 91:943–949
Pharoah PD, Dunning AM, Ponder BA, Easton DF (2004) Association studies for finding cancer-susceptibility genetic variants. Nat Rev Cancer 4:850–860
Mavaddat N, Pharoah PD, Michailidou K et al (2015) Prediction of breast cancer risk based on profiling with common genetic variants. J Natl Cancer Inst 107:djv036
Warren Andersen S, Trentham-Dietz A, Gangnon RE et al (2013) The associations between a polygenic score, reproductive and menstrual risk factors and breast cancer risk. Breast Cancer Res Treat 140:427–434
Reeves GK, Travis RC, Green J et al (2010) Incidence of breast cancer and its subtypes in relation to individual and multiple low-penetrance genetic susceptibility loci. JAMA 304:426–434
Hsieh YC, Hung CT, Lien LM et al (2009) A significant decrease in blood pressure through a family-based nutrition health education programme among community residents in Taiwan. Public Health Nutr 12(4):570–577
Long J, Cai Q, Shu XO et al (2010) Identification of a functional genetic variant at 16q12.1 for breast cancer risk: results from the Asia Breast Cancer Consortium. PLoS Genet 6:e1001002
Hein R, Maranian M, Hopper JL et al (2012) Comparison of 6q25 breast cancer hits from Asian and European Genome Wide Association Studies in the Breast Cancer Association Consortium (BCAC). PLoS ONE 7:e42380
Shu XO, Long J, Lu W et al (2012) Novel genetic markers of breast cancer survival identified by a genome-wide association study. Cancer Res 72:1182–1189
Zheng W, Wen W, Gao YT et al (2010) Genetic and clinical predictors for breast cancer risk assessment and stratification among Chinese women. J Natl Cancer Inst 102(13):972–981
Zheng W, Zhang B, Cai Q et al (2013) Common genetic determinants of breast-cancer risk in East Asian women: a collaborative study of 23 637 breast cancer cases and 25 579 controls. Hum Mol Genet 22(12):2539–2550
Steyerberg EW, Harrell FE Jr, Borsboom GJ et al (2001) Internal validation of predictive models: efficiency of some procedures for logistic regression analysis. J Clin Epidemiol 54(8):774–781
Efron B, Tibshirani R (1997) Improvements on cross-validation: the 632+ bootstrap method. J Am Stat Assoc 92:548–560
Harlid S, Ivarsson MI, Butt S et al (2012) Combined effect of low-penetrant SNPs on breast cancer risk. Br J Cancer 106:389–396
Sueta A, Ito H, Kawase T et al (2012) A genetic risk predictor for breast cancer using a combination of low-penetrance polymorphisms in a Japanese population. Breast Cancer Res Treat 132:711–721
Huang CS, Lin CH, Lu YS, Shen CY (2010) Unique features of breast cancer in Asian women–breast cancer in Taiwan as an example. J Steroid Biochem Mol Biol 118:300–303
Turner N, Grose R (2010) Fibroblast growth factor signalling: from development to cancer. Nat Rev Cancer 10:116–129
Cui F, Wu D, Wang W, He X, Wang M (2016) Variants of FGFR2 and their associations with breast cancer risk: a HUGE systematic review and meta-analysis. Breast Cancer Res Treat 155:313–335
Klinge CM, Blankenship KA, Risinger KE et al (2005) Resveratrol and estradiol rapidly activate MAPK signaling through estrogen receptors alpha and beta in endothelial cells. J Biol Chem 280:7460–7468
Zheng Q, Ye J, Wu H, Yu Q, Cao J (2014) Association between mitogen-activated protein kinase kinase kinase 1 polymorphisms and breast cancer susceptibility: a meta-analysis of 20 case-control studies. PLoS ONE 9:e90771
Yu Y, Chen Z, Wang H, Zhang Y (2013) Quantitative assessment of common genetic variants on chromosome 5p12 and hormone receptor status with breast cancer risk. PLoS ONE 8:e72154
Liang H, Li H, Yang X et al (2016) Associations of genetic variants at nongenic susceptibility loci with breast cancer risk and heterogeneity by tumor subtype in Southern Han Chinese Women. Biomed Res Int 2016:3065493
Han W, Woo JH, Yu JH et al (2011) Common genetic variants associated with breast cancer in Korean women and differential susceptibility according to intrinsic subtype. Cancer Epidemiol Biomark Prev 20:793–798
Long J, Shu XO, Cai Q et al (2010) Evaluation of breast cancer susceptibility loci in Chinese women. Cancer Epidemiol Biomark Prev 19:2357–2365
Wadt KA, Aoude LG, Golmard L et al (2015) Germline RAD51B truncating mutation in a family with cutaneous melanoma. Fam Cancer 14:337–340
Chan M, Ji SM, Liaw CS et al (2012) Association of common genetic variants with breast cancer risk and clinicopathological characteristics in a Chinese population. Breast Cancer Res Treat 136:209–220
Wacholder S, Hartge P, Prentice R et al (2010) Performance of common genetic variants in breast-cancer risk models. N Engl J Med 362(11):986–993
Hüsing A, Canzian F, Beckmann L et al (2012) Prediction of breast cancer risk by genetic risk factors, overall and by hormone receptor status. J Med Genet 49:601–608
Acknowledgements
This study was supported by the Health and Welfare Surcharge of Tobacco Products, Ministry of Health and Welfare to the Comprehensive Cancer Center of Taipei Medical University (MOHW105-TDU-B-212-134001), Taipei Medical University Hospital (103TMU-TMUH-09), Taipei Medical University (TMU102-AE1-B03), and the Top University Project- Cancer Translational Center of Taipei Medical University.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Chin-Sheng Hung and Hung-Yi Chiou contributed equally to this study.
Rights and permissions
About this article
Cite this article
Hsieh, YC., Tu, SH., Su, CT. et al. A polygenic risk score for breast cancer risk in a Taiwanese population. Breast Cancer Res Treat 163, 131–138 (2017). https://doi.org/10.1007/s10549-017-4144-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-017-4144-5