Abstract
Biological membrane remodeling is central to living systems. In spite of serving as “containers” of whole-living systems and functioning as dynamic compartments within living systems, biological membranes still find a “blue collar” treatment compared to the “white collar” nucleic acids and proteins in biology. This may be attributable to the fact that scientific literature on biological membrane remodeling is only 50 years old compared to ~ 150 years of literature on proteins and a little less than 100 years on nucleic acids. However, recently, evidence for symbiotic origins of eukaryotic cells from data only on biological membranes was reported. This, coupled with appreciation of reproducible amphiphilic self-assemblies in aqueous environments (mimicking replication), has already initiated discussions on origins of life beyond nucleic acids and proteins. This work presents a comprehensive compilation and meta-analyses of data on self-assembly and vesicular transformations in biological membranes—starting from model membranes to establishment of Influenza Hemagglutinin-mediated membrane fusion as a prototypical remodeling system to a thorough comparison between enveloped mammalian viruses and cellular vesicles. We show that viral membrane fusion proteins, in addition to obeying “stoichiometry-driven protein folding”, have tighter compositional constraints on their amino acid occurrences than general-structured proteins, regardless of type/class. From the perspective of vesicular assemblies and biological membrane remodeling (with and without proteins) we find that cellular vesicles are quite different from viruses. Finally, we propose that in addition to pre-existing thermodynamic frameworks, kinetic considerations in de novo formation of metastable membrane structures with available “third-party” constituents (including proteins) were not only crucial for origins of life but also continue to offer morphological replication and/or functional mechanisms in modern life forms, independent of the central dogma.
Graphic Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The discovery of enzymes as cellular constituents was a major landmark in the history of science (Kühne 1877). Prior to this discovery, appreciation of “small” single cells as living units, having the ability to appear de novo, was also coupled with the belief that only living cells (called ferments) had a “magical” ability to carry out a variety of chemical transformations at ambient conditions. This “magic” was a signature of life. The fact that all enzymes were proteins (believed for more than a century after first use of the word “enzyme” by Kühne) immediately put the onus of biological replication (and evolution of living systems since the origins of life) on proteins. These were molecules found in all living cells and, now known to be responsible for all biochemical reactions. More than half-a-century later, this onus shifted to DNA subsequent to remarkable work of Griffith (1928), Oswald et al. (1944), Chargaff (1950), Hershey and Chase (1952), Rosalind Franklin, and Maurice Wilkins, that led to often called “discovery-of-the-century”, i.e., structure of DNA by Watson and Crick (1953). Almost another half-a-century later, discovery of catalytic functions of RNA by Altman and Cech (Kruger et al. 1982; Guerrier-Takada et al. 1983) initiated discussions on an RNA-world hypothesis for origins of life (Cech 2012). In a span of ~ 100 years covered above, views on molecular basis of origins of life, biological replication, and evolution of biological systems shifted from proteins to DNA to possibly DNA and RNA. Emerging from this time span, functional manifestations of proteins now define “phenotypes” and nucleic acids are considered as indicators of biological relatedness in experimental as well as evolutionary time scales (Woese et al. 1990). All this while, biological membranes (specifically their lipid constituents), in spite of being the “third front” in living systems, i.e., beyond nucleic acids and proteins, continue to remain as comparatively passive components in origins of life and biological evolution (Bansal and Mittal 2015). In fact, a common “textbook” visualization of cell membranes comes from Singer and Nicolson (1972)—the fluid-mosaic model in which a mosaic of non-phospholipid entities (e.g., integral proteins) is embedded in a phospholipid bilayer acting as two-dimensional viscous fluid. More than a decade later, a phenomenological thermodynamic perspective with a purely mechanical emphasis resulted in a mattress model (Mouritsen and Bloom 1984)—a cell membrane was visualized as an elastic mattress of lipids with non-lipid impurities (e.g., proteins) embedded as localized springs (with distinct elasticities) in the mattress. While neither of the above models focused on heterogeneous details (e.g., local curvatures), they allowed visualization of “average” cell membranes through specific organizational frameworks; the former providing some features relevant to nano-scales while laying foundations for the well-appreciated asymmetry in cell membranes. Over the years, these models have transformed into a more mosaic view of cell membranes by incorporating localized, often transient, interactions (including those involving peripheral membrane and cytoskeletal proteins along with components of extracellular matrix) resulting in distributions of heterogeneous domains/rafts with different relative mobilities (Jacobson et al. 1995; Nicolson 2013; Almeida 2014; Nicolson and Ferreira de Mattos 2021). However, the above visualizations of cell membranes, while increasingly attributing active roles to their lipid components, still do not consider those lipids in contexts of origins of life and biological replication (Mittal et al. 2020).
Some of the earliest work on development of model membrane systems (Bangham and Horne 1964; Bangham et al. 1965), transitioning into development of liposomes (Bangham 1972), indicated close similarities to behavior of biological membranes—on the level of whole cells as well as intracellular organelles. In fact, the early excitement of being able to replicate both biological structures and their behavior in absence of proteins and nucleic acids is well captured by the following representative statement from Bangham and Horne (1964): Certainly the lesions produced in the model membrane of lecithin–cholesterol are commensurate with the damage obtained in biological cells when either saponin or lysolecithin is used as a lytic agent. In spite of the above and a sustained interest in the possibilities of creating “simple” protocells and life-like properties from “non-living” materials (Rasmussen et al. 2004; Mittal 2009), biological replication as a “property” still resides only in the domain of the central dogma. In this context, it needs to be appreciated that literature on experimental kinetics and thermodynamics of membrane remodeling, starting from protein-free systems to protein-mediated transitions/transformations, is only ~ 50 years old. In contrast, there are over 100 years of literature on proteins and a little less than 100 years of literature on nucleic acids. Therefore, it is not surprising that exploration of possible biological replication mechanism(s) offered by self-assembling lipid systems in aqueous environments, free from nucleic acids and proteins, is very recent (Mittal et al. 2020; Steinkühler et al. 2020, 2021). However, proper analyses of scientific literature on remodeling in self-assembled membranes and/or vesicular transformations opens up possibilities of “phenotypic” and/or “functional” replication mechanisms in nucleic acid- and protein-free aqueous environments. Further, transitional events recorded during protein-mediated membrane remodeling show availability of kinetic windows where thermodynamic constraints may become available for (re)creation of structures and/or functions in protein-free systems. In this regard, one of the most biologically relevant and ubiquitous membrane remodeling events is membrane fusion. From the variety of structural and transformational aspects involved in fusion of bilayers, to kinetic and thermodynamic considerations during morphological transitions, membrane fusion covers almost all “variables” pertaining to membrane remodeling.
In this work, we comprehensively analyze literature on assembly and transformations in biological membranes, starting from protein-free vesicular transformations and culminating in protein-mediated membrane remodeling, with a focus on membrane fusion. To every extent possible, we have collected data from original experimental literature (that has withstood the test of time and scrutiny) instead of relying on specific interpretations of that data presented in numerous reviews. While highlighting and appreciating some landmark experimental results on kinetics and thermodynamics of protein-free vesicular systems, we discuss the emergence/establishment of Influenza Hemagglutinin (HA, also spelled as Haemagglutinin) as a prototypical membrane fusion protein. In fact, HA is arguably as important for understanding molecular-level functioning in biology from the perspective of ubiquitous membrane remodeling dynamics crucial for life, as have been enzyme systems from the perspective of the protein folding problem. We discuss how HA-mediated membrane fusion systems have attained significance in biology similar to that of enzymes that provided “functional assays” (e.g., ribonuclease and chymotrypsin) and were responsible for opening the gates of current understanding on structure and function of proteins (Anfinsen 1972). Next, we make a comprehensive comparison of various biological vesicular systems—enveloped mammalian viruses, extracellular, and intracellular vesicles, especially based on size and certain physico–chemical properties in context of biological membrane remodeling. Finally, we lay the foundations of development of a theory on how kinetic transitions in membranes may be “stabilized” by proteins toward formation of dynamic, yet metastable, structures away from equilibrium configurations preferred by the same membrane constituents in aqueous systems. Such structures play integral roles in living systems and understanding their formation is crucial toward gaining insights into origins of life, biological self-assembly, and replication.
Protein-Free Vesicular Assemblies, Transformations, and Membrane Remodeling
Table 1 compiles sizes of vesicular assemblies (and transformations) formed (and reported) in some pivotal experimental studies with specific lipid compositions in aqueous systems. Each of these studies represent significant methodological and/or analytical advances, including experimental assays and their interpretations. While the pre-1975 studies mostly relied on freeze fracture and electron microscopy techniques for visualization of membranes assemblies and vesicular transformations, phase behaviors of lipid mixtures were quantitated using differential scanning calorimetry or spectra of spin labels distributions that depended on membrane phases. By 1980, protein-free systems had developed enough to be able to provide interesting insights into “functional” aspects of biological membranes, such as permeability, in addition to visible mimicking of compartmental morphologies observed in cells. Subsequent to refinements in methods of liposomal preparations, 1980 to 1990 was a remarkable decade resulting in development of crucial membrane fusion assays for varying liposome sizes (size represents curvature, especially in smaller assemblies). Contents mixing assays using Tb3+/dipicolinic acid, carboxyfluorescein, drug molecules, enzymatic substrates, and a variety of different sized molecules transferring between fusing compartments were developed. In addition, lipid mixing assays (not listed in Table 1), namely—dequenching of the self-quenched R18 (Octadecyl Rhodamine B) upon lipid mixing (Struck et al. 1981), FRET between N-(7-nitro-2,1,3-benzoxadiazol-4-yl)—Rhodamine subsequent to lipid mixing between bilayers separately labeled with NBD-PE and N-Rh-PE (Hoekstra et al. 1984) and the Pyrene excimer assay (Stegmann et al. 1993) were also established. The Pyrene assay overcame some of the limitations of the earlier lipid mixing assays such as non-specific probe transfer on close apposition of bilayers. A key achievement in this period was the development of rigorous kinetic frameworks (e.g., Bentz et al. 1983a, 1983b) and thermodynamic constraints (e.g., Lentz et al. 1987) required to properly interpret data on vesicular preparations and their utilization in membrane remodeling experiments. Analytical tools clearly extracting distinct kinetics of aggregation, lipid mixing, contents mixing, and final extents of measurements (fusion) from the data were developed. Specifically noted was the fact that neither were vesicular assemblies at thermodynamic equilibrium, nor did the end points of assays depict thermodynamic end points of membrane remodeling. These rigorous quantitative treatments of experimental data on fusion of protein-free lipid bilayers (and later protein-mediated membrane fusion) laid the foundations for a variety of subsequent studies on protein-mediated remodeling of biological membranes directly or indirectly, including the Nobel-accorded (in 2013) efforts on discoveries of machinery regulating vesicle traffic, a major transport system in our cells. (Malhotra et al. 1988; Perin et al. 1990; Sollner et al. 1993; Hata et al. 1993; Weber et al. 1998).
By early 1990s, developments in (and more accessibility of) fluorescence microscopy (epifluorescence followed by confocal) allowed direct visualization of vesicular assemblies, phase separations, and dynamics of membrane remodeling. Thus, as is evident from Table 1, substantial contributions emerged out of work on preparation and visualization of GUVs. This allowed experimental verification, and further development, of concepts such as line tension, mean/Gaussian curvatures, bending rigidity, and tilt/splay in lipids that are predominantly theoretical (but physically important). Start of the new century brought progress toward quantitative visualization of transitions and distributions of different phases in individual membrane assemblies compared to populations of assemblies. Important phase diagrams of defined amphipathic mixtures self-assembling into membranes in aqueous environments continue to emerge out of such progress till date. Interestingly, the last couple of entries in Table 1 represent a somewhat cyclical nature of scientific exploration—(i) the work of Steinkühler et al. (2018) is reminiscent of the seminal work of Chernomordik et al. (1987) that investigated different stages of electric field-driven morphological transitions and curvature formations in planar bilayers and (ii) the work of Spustova et al. (2021) is reminiscent of the seminal work of Wilschut et al. (1980), Nir et al. (1982), and Bentz et al. (1983b); spontaneous vesiculation and “subcompartment” formation on removing Ca2+ observed recently may be simply interpretable as the dominance of reverse rate constants in the older studies that used Ca2+ as an inducer for creation of larger assemblies (fusion products) from smaller vesicles. Nevertheless, the recent studies allow a much better appreciation of the original literature from 1980s, especially for interpreting experimental results on protein-free vesicular assemblies, their transformations, and overall membrane remodeling in the context of origins of life and structural replication mechanisms (of biological compartments) independent of the central dogma.
In addition to the above, some very recent methodological advances provide fascinating ways to address a key limitation of previous vesicular assemblies. Unlike cell membranes, liposomal membranes are devoid of membrane asymmetry. Bhatia et al. (2018) overcame this limitation by inducing (transient) asymmetry in GUV bilayers by utilizing small amounts of glycolipids (ganglioside) GM1 whose local concentrations in respective monolayers changed due to desorption of GM1 from the outer monolayer as a result of extravesicular “dilution” by preparation buffer. Subsequently, Bhatia et al. (2020) induced (transient) asymmetry in osmotically balanced GUVs with encapsulation of a sugar different from the extravesicular sugar in the suspension solution, thereby varying local hydration environments of the inner and outer monolayers. Such innovative approaches resulting in remarkable protein-free vesicular assemblies and transformations are gradually pushing “in vitro” systems closer to mimicking previously “unmatched” features of living cells (“in vivo” systems).
From Advent of Liposomes to Development of Experimental Assays for Membrane Fusion
Figure 1 shows a timeline from the advent of liposomes to development of experimental assays for membrane fusion; initially in protein-free and later applied to protein-mediated membrane fusion experiments. By early 1970s, model membrane systems had been developed from a variety of amphipathic mixtures in different aqueous environments (Bangham 1972). While experimental work on “functional” manifestations (e.g., permeability, rupture/repair) of these model membranes continued, theoretical frameworks and mechanistic models involving physico–chemical properties of amphipathic molecules in aqueous systems emerged. Today, conceptual formulations of (i) the hydrophobic effect (Tanford 1973; Tanford 1978; see legend to Fig. 1) and, (ii) shape parameters for amphipathic molecular constituents (Israelachvili et al. 1976) of assemblies in aqueous environments (e.g., micelles, bilayers), have the same significance in understanding of biological membranes as secondary structures are significant in understanding of protein structures. In fact, such is the remarkable validity and applicability of a theoretical concept such as shape parameter that it is found to be a molecular signature of bilayer constituents even in simulations of flat bilayers (Bansal and Mittal 2013).
Timeline from advent of liposomes to development of experimental systems and assays for membrane assemblies and fusion (see text for details). The timeline captures some of the key original experimental advances—from the first liposomes in the 1960s, to the latest protein-free vesicular assemblies involving “electroformation” of GUVs from heterogenous SUVs (Bhatia et al. 2015) along with inducing of transient bilayer asymmetries by varying intra- and extra- vesicular conditions (Bhatia et al. 2018, 2020). Here, it is pertinent to enforce the importance of properly interpreting the hydrophobic effect—the term “hydrophobic interaction” neither implies any force nor is it a real “interaction”—it is actually exclusion by water. In absence of an aqueous environment, “hydrophobic” molecular entities do not show any tendency to interact with each other. Thus, while the word “hydrophobic” may appear to imply phobia from water, experimentally and scientifically it represents only exclusion from/by liquid water (anecdotally interpretable as water’s phobia for such molecules/entities rather than the other way around). The figure is drawn only for illustration purposes and not to any scale
The 1980s also saw complementation of membrane fusion between curved membranes by visualization of flat planar bilayers spontaneously formed by defined compositions in aqueous environments with the help of Teflon and similar materials serving as substrates/surfaces. Chernomordik et al. (1987) utilized/developed one such experimental assembly for recording structural transitions during membrane fusion between two planar bilayers while coupling their experimental findings with a theoretical stalk model (Kozlov and Markin 1983). In fact, on one hand, the work of Wilschut et al. (1980), Bentz and Nir (1981a, b) created the foundations for combining theoretical and experimental kinetics in membrane fusion. On the other hand, Chernomordik et al. (1987), Ellens et al. (1989) and Siegel (2008) created the foundations for combining theoretical and experimental thermodynamics of remodeling during membrane fusion. Next, as introduced earlier, developments in fluorescence microscopy allowed direct and visual measurements of membrane fusion. In this direction, Blumenthal, Zimmerberg and colleagues, utilized erythrocytes [red blood cells (RBCs)] and RBC “ghosts” fusing with HA-expressing cells for direct monitoring of events occurring during biological membrane fusion (Morris et al. 1989; Sarkar et al. 1989; Herrmann et al. 1993). This was possible since glycophorins on RBCs are heavily sialated and sialates serve as receptors for HA. The above was made possible by development of the HA-expressing cell—RBC ghost experimental system by Doxsey et al. (1985) and Sambrook et al. (1985). With this, the era of studies on membrane fusion of protein-free lipid bilayers transitioned to predominantly protein-mediated membrane fusion. The basic fundamentals of experimental assays pioneered during that time continue to serve till date, with liposomes (especially SUVs representing curvature-related parameters in bilayers, and LUVs) and protein-free lipid systems (“lipid nanosystems”) shifting in relevance as drug delivery vehicles in subsequent times (Meers 2022).
In relation to the above, it is pertinent to note that later applications involving X-ray (diffraction, scattering) and/or neutron scattering have allowed a high-resolution structural characterization of protein-free lipid bilayers further validating and/or confirming earlier findings. While predominantly using flat model membranes, these experiments have resulted in quantitative estimates for bilayer interaction forces (Wong et al. 1999a, b) along with properties such as membrane thickness, degree of heterogeneity, tilt, bending moduli and phase separations etc. (Karmakar and Raghunathan 2005; Kamal and Raghunathan 2012; Tristam-Nagle 2015; Nagle 2017). Specific high-resolution visualizations of spatial organizations and/or local curvatures and/or suitability of membrane systems for (or resistant to) remodeling continue to complement/confirm earlier results obtained from relatively low-resolution assays and observations (see lower left panels “2002-till date” in Fig. 1). For example, on one hand Yang and Huang (2002, 2003) were able to directly visualize a “static” stalk formation in dehydrating lipid membrane stacks. On the other hand, Salditt and colleagues captured transient “dynamic” stalk formations, presenting them as structural fusion assays, in aligned multi-lamellar stacks of protein-free-lipid membranes (Aeffner et al. 2012; Scheu et al. 2021).
Influenza HA-Mediated Membrane Fusion: Establishment of a Prototypical Fusion System
In the late 1960s, first reports pertaining to internalization of several viruses (Type 2 Parainfluenza virus, Influenza virus, New Castle Disease virus, Herpes Simplex virus, and Sendai virus) by different cells came from light microscopy (Howe et al. 1967) and electron microscopy (Howe et al. 1967; Morgan and Howe 1968 and references there in). These studies did indicate events such as “binding,” “fusion,” “penetration,” and “uncoating” of virus particles with different cells—based on protocols involving incubation of virus particles at 4 °C for “binding” and raising the temperature to 37 °C for “penetration” into cells. Using a similar protocol and electron microscopy, Haywood (1974) reported “fusion” and “penetration” of Sendai virus into ganglioside containing model membranes, presumably to avoid cell culture-related artifacts. However, artifacts pertaining to sample preparation and/or image acquisition in all the above experiments were still difficult to segregate. It is interesting to note that Sendai virus, known to induce syncytia in cell cultures, does not require low pH for membrane fusion. Influenza virus did not induce similar syncytia but was shown to “penetrate” target membranes in the above studies. Clearly something was amiss in experimental protocols. To this end, Helenius et al. (1980) followed by White and Helenius (1980) reported low pH-driven membrane fusion by the Semliki Forest virus (SFV), using assays involving cells and liposomes, but free of possible artifacts in the earlier studies. The fusion assays were “end-point” and comparatively robust. Catalytic activity of enzyme trapped in liposomes on viral RNA subsequent to fusion and inhibition of viral infectivity in cells treated with lysosomotropic agents (that increased lysosomal pH) were indirect assays compared to direct kinetic assays (Table 1). However, the experiments were ground-breaking for discovering the need for low pH “trigger” required for SFV membrane fusion. The aftermath was unequivocal demonstration of low pH-induced membrane fusion by Influenza and the Vesicular Stomatitis viruses—the experimental assay was formation of cell syncytia (White et al. 1981).
Figure 2 shows the timeline for establishment of Influenza HA as a prototypical membrane fusion protein (note that the figure also includes crucial pre-influenza HA experiments also for completeness). Ectodomain of HA was the first membrane fusion protein whose crystal structure was solved (Wilson et al. 1981). The solved structure was that of bromelain-cleaved HA at neutral pH and the remaining decade (plus some more years) went into consistent efforts to decipher low pH conformation(s) of HA. A key development was creation of HA-expressing fibroblast cell lines (Doxsey et al. 1985; Sambrook et al. 1985) that allowed development of HA-mediated cell–cell membrane fusion systems triggered by low pH. This resulted in some exemplary experiments monitoring double-labeled RBC ghosts fusing with HA-expressing cells when exposed to low pH. Experimental protocols developed for model membrane fusion systems were now directly applied to HA-mediated fusion. Convenient size of the cell–cell fusion system allowed real-time monitoring of fusion events using patch clamp, lipid mixing, and contents mixing assays. Influential discoveries on sequence of events, from flickering pores allowing ion transfer, to lipid mixing, to opening, and expansion of fusion pores during protein-mediated membrane fusion, were made (Sarkar et al. 1989; Spruce et al. 1989; Tse et al. 1993; Zimmerberg et al. 1994; Leikina et al. 2004). These were accompanied by periodic experiments of Influenza virus fusing with liposomes (e.g., Stegmann et al. 1995). However some key questions remained: what was(were) the exact role(s) of HA in fusion? What was the architecture of the fusion site? These questions were reflective of almost a century-old investigation on how proteins work as enzymes (see “Introduction” section).
Timeline for establishment of Influenza HA as a prototypical membrane fusion protein and development of experimental assays for studying HA-mediated membrane fusion (see text for details). Wilson et al. (1981) solved the structure of bromelain-cleaved HA (called BHA) at neutral pH—the first structure shown is from PDB (Berman et al. 2000) PDB ID: 5HMG (Wilson et al. 1981; Weis et al. 1990); the second structure, called TBHA2, shown is PDB ID: 1HTM (Bullough et al. 1994) which is trypsin + thermolysin + bromelain—cleaved HA at low pH. Overall interpretation from 5HMG and 1HTM is that HA is a trimer with each monomer having two subunits—HA1 responsible for receptor binding and HA2 responsible for membrane fusion. The figure is drawn only for illustration purposes and not to any scale
To this end, Ellens et al. (1990) conducted one of the most seminal experiments, which introduced experimental transformation of fundamental concepts of three-dimensional chemical reaction stoichiometry from solutions to lesser dimensions (in this case surface reactions). Monitoring fusion of fibroblasts expressing different surface densities of HA to same target membranes was equivalent to laboratory titrations aimed at uncovering chemical reaction stoichiometries in solutions. And the results were astounding—HA surface density was not linearly related to the extents of fusion observed. Ellens et al. (1990) unambiguously showed involvement of more than one HA in fusing membranes and allowed development of the first estimates of the architecture of a protein-mediated membrane fusion site (Bentz et al. 1990). However, a more accurate estimation of the fusion site architecture required kinetic data in addition to extents. Further, assignment of specific roles to individual HA molecules in the fusion site required knowledge of low pH structure of HA. In regards to the latter, a remarkable computationally driven prediction by Carr and Kim (1993) was confirmed by solving of low pH crystal structure of HA, called TBHA2 (shown at 1994 in Fig. 2, explained in figure legend). The structure not only showed one of the most dramatic conformational changes discovered in any protein till that date, but also provided possible mechanisms for membrane fusion along with an experimentally verifiable concept of “metastable” states in general protein folding. This was followed by a series of experiments that generated invaluable kinetic data on HA-mediated membrane fusion with a variety of experimental systems—first fusion pore formation in cell-planar bilayer fusion (Melikyan et al. 1995), virus-liposome fusion at different pH values, with different strains of Influenza with different target membranes (Shangguan et al. 1996), and cell–cell fusion (Danieli et al. 1996; Blumenthal et al. 1996). Kinetic data generated above was comprehensive since (a) it used the same standardized HA-expressing cell lines or quantitatively well-characterized Influenza virus particles and (b) it came from diverse, but equally important methods, for monitoring different steps of membrane remodeling during fusion till that date, i.e., electrophysiological measurements, lipid mixing, and contents mixing assays. Interestingly, individual interpretations of the data did not appear to converge on the same (or even similar) architecture of the fusion site - differences attributable to different measurements (single-first-fusion-pore-event vs single-cell–cell-fusion-membrane-conductivity vs single-cell–cell–lipid/content mixing vs population-cell–cell–lipid-mixing vs population-virus–liposome–lipid/content mixing).
Remarkably, a mass-action model initially developed by Bentz (2000a), extended by Mittal and Bentz (2001), followed by Mittal et al. (2002b), was able to extract an unambiguous consensus architecture of the HA-mediated membrane fusion site from the above comprehensive kinetic datasets (Bentz and Mittal 2003). Specific roles were assigned to receptor-bound HA and free HA at the fusion site while incorporating cryo-EM based data of Shangguan et al. (1998) on low pH induced inactivation (i.e., fusion incompetence) of influenza virions. The emergent “textbook” picture defined a minimal aggregate size (ω = minimum number of HA molecules aggregated to form a fusion site) and a minimal fusion unit (n = minimum HA molecules required to undergo the dramatic conformational changes observed from BHA to TBHA2 for creation of the first fusion pore). Based on the data, receptor-bound HA molecules were not a part of the “n” (minimal fusion unit) and assisted in creation of restricted fusion site that eventually expands. The significance of this mass-action model to HA-mediated membrane remodeling during fusion is arguably akin to that of Michaelis–Menten kinetics in enzyme function—with an additional advantage of the primary parameters in the fusion model, i.e., ω and n, being mechanistic (in contrast to VMax and KM which are phenomenological). While the above was focused on arriving at the architecture of HA-mediated membrane fusion site from the perspective of HA molecules, Chernomordik et al. (1997, 1998) carefully dissected lipidic intermediates formed during HA-mediated merger of bilayers using cell–cell fusion assays—with fascinating approaches that involved arresting or promoting membrane curvature-based remodeling during HA-mediated fusion by addition of exogenous lipids with different shape parameters to the experimental system. By combining electrophysiological measurements, lipid, and contents mixing assays, Chernomordik and colleagues were able to create “textbook” visualizations for protein-mediated membrane remodeling events during HA-mediated membrane fusion. This concluded a comprehensive series of analytical and experimental efforts with identification of distinctions between HA-mediated membrane remodeling during hemifusion and fusion of bilayers (Mittal et al. 2003). Importantly, apart from establishing Influenza HA-mediated membrane fusion as a prototypical system, the analytical and experimental advances described in this section (and shown in Fig. 2) continue to serve as reliable tools in understanding protein-mediated remodeling during membrane fusion in general, till date.
From Influenza HA-Mediated Membrane Fusion to Other Enveloped Mammalian Viruses
Table 2 lists the morphological and size variations in enveloped mammalian viruses, along with respective viral membrane proteins (VMPs, with subunits wherever applicable) responsible for receptor binding to the host and/or membrane fusion with the host. Here, an important point to note is that similar to classifications of general proteins into alpha, beta, and alpha/beta classes (Mittal and Acharya 2012; 2013), VMPs are also classified (into Classes I, II, III) based on alpha-helical and beta-sheet contents (White et al. 2008; Backovic and Jardetzky 2009; Modis 2014). Here, we do not focus on those classifications. However, we list several common themes emerging from HA-mediated membrane remodeling, which are applicable to all of viruses listed in Table 2:
- 1.
Lipidic intermediates in protein-mediated membrane remodeling during fusion by enveloped viruses are the same (though with kinetic variations), regardless of structural variations in VMPs (Zaitseva et al. 2005).
- 2.
All VMPs, without exception, have a “fusion peptide”—an amphipathic/hydrophobic stretch of amino acids that plays a key role in destabilizing viral and/or target membranes. Gething et al. (1986) first established the importance of this peptide by demonstrating that even a single-point mutation in HA fusion peptide renders the whole protein ineffective in membrane fusion. Length of this fusion peptide can be 10–25 residues (Bentz and Mittal 2000) and it is generally hidden/buried in the “soluble” ectodomain of VMPs before fusion. Exposure of this fusion peptide is a result of conformational changes in response to some trigger (e.g., receptor binding of VMP or lowering of pH).
- 3.
Metastable conformations of VMPs—initial conformations of VMPs undergo irreversible conformational changes (i.e., final conformations are more “stable”) resulting in membrane fusion. The paradigm of natively occurring less stable conformations in natural proteins, emerging from HA, is in contrast to expectations based on findings of Anfinsen (1972). Thus, the concept of “metastability” in the initial conformations of VMPs appropriately reconciles the post-fusion conformational irreversibility with Anfinsen’s thermodynamic view. This also results in “activation” and “inactivation” of VMPs—Priming for irreversible conformational changes by triggers (e.g., low pH or receptor binding) in presence of target membranes leads to membrane fusion and in absence of target membranes inactivates virus particles. Till date, only rabies virus G protein has emerged as an exception—it exhibits reversible conformational changes and can catalyze multiple rounds of fusion (Gaudin et al. 1991).
- 4.
Occurrence of coiled coils or “n”-helix-bundles or hairpins in VMPs—these structural/conformational motifs are either present in pre-fusion metastable conformations or in many cases are a result of conformational changes in VMPs coupled with pre- to post-membrane fusion events (Bentz 2000b). These conformational motifs are often credited for ensuring close apposition of membranes (overcoming hydration barriers and/or generating local curvatures at fusion sites to facilitate membrane fusion).
- 5.
Fusogenic aggregates/Fusion machines or units—multiple VMP molecules assemble together to create the required architecture of a membrane fusion site. Incomplete or different VMP assemblies lead to hemifusion or no fusion.
- 6.
Leaky fusion—protein-mediated membrane fusion can be leaky, but only for very small molecules.
- 7.
Data obtained from population assays for membrane fusion can be directly mapped to single-fusion events or single cell–cell fusion measurements (Mittal et al. 2002a). Interestingly, for several other systems in biology, such as single-molecule experiments, this still remains a challenge.
- 8.
Local membrane environments (e.g., cholesterol enrichment or specific membrane domains) around VMPs are important in modulating their membrane fusion activities (Hess et al. 2005, 2007; Biswas et al. 2008; Yang et al. 2015; Lee et al. 2021).
In addition to (and preceding the above), VMPs play crucial role(s) in close apposition of viral membranes with target membranes—this close apposition requires compensating for, followed by overcoming of, the hydration barriers between the outer monolayers of the two fusing bilayers. While being a general aspect of biological membrane fusion, this involves local dehydration coupled with possible creation of transient hydrophobic defects required to be “healed” by exchange (or flip-flop) of outer monolayer lipids and/or protein fragments (Tieleman and Bentz 2002; Witkowska et al. 2021). Here, it is also important to highlight that, other than viral membrane fusion, many of the above themes are also common to ubiquitous protein-mediated membrane remodeling events in cellular and physiological systems, such as embryonic development in worms (Gattegno et al. 2007).
Having discussed common themes on viral fusion mechanisms above, it also pertinent to view fusogenic components of VMPs (e.g., HA2 in HA) as general proteins, instead of a restricted view as only fusion proteins. Recently, it emerged that naturally occurring folded/structured proteins have clear compositional constraints (Mittal et al. 2010, 2020; Mittal and Jayaram 2011a, b). It was also shown that amino acid compositions beyond those constraints are signs of intrinsic disorder in proteins, i.e., lack of specific conformations/structures corresponding to functions (Mittal et al. 2021a, b, c). Thus, considering structural classifications in fusogenic components of VMPs (White et al. 2008), it was natural to test whether VMPs obey “stoichiometry-driven protein folding” (Agutter 2011). Figure 3 shows that VMPs are highly structured (black and yellow bars compared to striped bars in Fig. 3A), regardless of whether they are predominantly alpha-helical or beta-sheets. Also, the variability in VMP compositions, in spite of primary sequences being very different, is extremely low especially when compared to intrinsically disordered proteins (Fig. 3B). These results not only re-iterate the crucial role of relative occurrence of amino acids in naturally occurring structured proteins but also show that VMPs are highly structured with even tighter compositional constraints than general-structured proteins.
Stoichiometric distributions of amino acids in viral fusion proteins (VMPs). These are compared with “structured” (open bars, n = 27,199), “sequences without structure” (gray bars, n = 532,553), “curated/reviewed intrinsically disordered proteins” (black-striped bars, n = 707), and “putative intrinsically disordered proteins” (gray-striped bars, n = 94)→for data and details, see Mittal et al. (2021c). The following sequences of fusogenic (components of) VMPs were collected from UniProtKB—HA2 (HA2-X31: P03437, HA2-Jap: P03451, HA2-PR8: P03452) and other viral fusion proteins (HIV1-gp41: P03375, HIV1-gp41: P03378, HIV2-gp41: P15831, HIV2-gp41: P20872, SFV-E1: Q8JMP5, Sin-E1:P03316, Sin-E1: P27285, TBE-E: P07720, TBE-E: P14336, TBE-E: Q01299, Den1-E: P27910, Den2-E: P29990). Yellow bars represent stoichiometric distributions of only HA2 (n = 3) and black bars represent stoichiometric distributions of all VMPs (n = 15)
Extra- and IntraCellular Vesicles are Different from Viruses
In the context of our discussions on protein-free membrane vesicles to enveloped mammalian viruses, it becomes important to inspect other naturally occurring vesicular systems in biology. Thus, considering that enveloped mammalian viruses are proteo-vesicles encapsulating cargo, we take a comparative look at extracellular and intracellular vesicles.
While there are online portals dedicated to extracellular vesicles, such as “Vesiclepedia”—www.microvesicles.org (Kalra et al. 2012; Pathan et al. 2019) and “Exocarta”—www.exocarta.org (Mathivanan et al. 2012; Keerthikumar et al. 2016), Table 3 here provides a succinct, straightforward, and yet comprehensive tabulation of broad/major types of extracellular vesicles (EVs) with information only specific to the current context. It is clear that in spite of an appealing parallel between “excretory” mechanisms releasing EVs and enveloped viruses, there is one physical difference between the two—size heterogeneity within individual viruses (Table 2) is much lower than the heterogeneity observed within individual EVs. Next, a physico–chemico–biological attribute unique to enveloped viruses is uniform spatial distributions of respective VMPs on viral envelopes/surfaces independent of their shapes as well as surface density. Moreover, the variety of VMPs for individual viruses are highly limited compared to possible types of different proteins associated with EVs. Finally, while functional roles of EVs are predominantly inter-cellular communications and/or transfer of materials between cells, viruses are known to primarily hijack intracellular machinery for their own replication. In view of the above, we find that extracellular vesicles are very different from viruses. Even if their cellular sources may appear similar, mechanisms of formation and the final vesicular forms are quite distinct. We propose that it may be important to consider viruses as signatures of very primitive vesicular forms (VFs), first arising out of self-assembly of purely amphipathic constituents in aqueous environments. These VFs later associated with proteins and nucleic acids (or vice versa) when these chemical species became available. Subsequently, the proteo-nucleic acid-VFs became a part of chemical hit and trials eventually leading to emergence of the central dogma. Here, it is important to emphasize that the above does not support any RNA-world hypothesis. Since DNA is a chemically more stable and less reactive molecule compared to RNA (Mittal 2012), DNA viruses simply could not evolve much compared to more reactive and thus prone-to-mutation RNA viruses. In fact, the dwindling variety of DNA viruses may compel viewing them as fossilized signatures supporting the origins of life from replication of VFs, developing into proteo-DNA-VFs→proteo-DNA/RNA-VFs independent of the central dogma, subsequently transforming into living cells emerging from the central dogma.
Table 4 shows a compilation of intracellular vesicles (IVs). Almost all the points discussed in comparing EVs with enveloped viruses are applicable to comparisons of IVs with the same viruses—except that functional roles of IVs are predominantly intracellular and/or for internalization of material from outside to inside cells. Since enveloped viruses utilize intracellular trafficking pathways, transmembrane domains (TMDs) of viral VMPs show a remarkable “length” signature for intracellular organelles involved in their internalization. Thus, based on distinct membrane environments for distinct organelles inside cells (Sharpe et al. 2010; Mittal and Singh 2018), internalization pathways of enveloped viruses can be predicted based on analyzing TMDs of VMPs (Singh and Mittal 2016). Here it is important to emphasize somewhat misleading aspects of “common features” of some IVs with viruses. Firstly, apparently both are vesicular forms with some sort of protein coats/assemblies associated with them. Secondly, already discussed previously, is the presence of common structural motifs in protein assemblies responsible for membrane fusion in both. These two need not be interpreted as similarities between IVs and enveloped viruses. The mechanisms of proteins’ association with IVs are very different than those in enveloped viruses, e.g., IV protein coats have proteins that have been shown as curvature sensors; due to functional implications on cellular dynamics, protein associations with IVs are through weak interactions. In contrast, VMPs are predominantly integral membrane proteins. Additionally, the structural motifs present in viruses arise from a very few types of proteins per virus (in many cases, it is a single type of protein) compared to IVs where multiple types of proteins come together via weak interactions to form these motifs which have non-uniform spatial distributions on vesicular surfaces.
Summarizing, close inspection of the data in Tables 2, 3, and 4 indicates that neither EVs nor IVs appear to have similar proteo-vesicle-cargo formation properties with enveloped viruses. It is also important to consider that cellular vesicles are compositionally “fragile,” not because of their membranes but because of reversible nature of weak interactions of proteins associated with them. In absence of data on half-lives of cellular vesicles in vivo and in vitro, a direct comparison of compositional “stability” with relatively more robust viral particles is not possible. Nevertheless, it is clear that any evolutionary linkages between viruses and cellular vesicles are premature at best and may even be biophysically unsound.
Discussion
The elegance and beauty of DNA structure offering a replication mechanism via the central dogma are undeniable. However, it is extremely difficult to visualize an accidental appearance of components of the central dogma in “dilute” solutions to initiate closely coordinated reactions for initiating life. Therefore, it is almost obvious/natural to envision appearance of replicable compartments that could encapsulate and constrain the components of central dogma to operate. Thus, origins of life must start with exploration of protein-free and nucleic acid-free replication mechanisms. These are seen in biological membranes. Recently, importance of lipid constituents of cellular “compartments” was explicitly demonstrated in origins of life and evolution of “complex” cells (Danchin 2014; Bansal and Mittal 2015). Comprehensive analyses of ~ 5000 lipid constituents of plasma membranes in the three domains of life provided direct, nucleic acids free, evidence for symbiotic origins of eukaryotic cells (Bansal and Mittal 2015). Also emerged was the reason for Archaea not being pathogens—membranes formed by sn-glycerol-1-phosphate do not fuse with membranes formed by glycerol-3-phosphate as subsequently noted by Antoine Danchin (personal communication). Clearly there is more to membrane remodeling by itself than protein-mediated membrane remodeling. In fact, it is important to consider the following:
- (1)
Membrane assemblies involve the same weak interactions as protein folding—these are the hydrophobic “interactions” (see legend to Fig. 1), hydrogen bonding, ionic interactions and Van der Waals forces. Of course, proteins also have two additional strong interactions (peptide bonds and disulfide linkages/bonds) that are absent in membranes—however, this absence allows morphological flexibilities.
- (2)
Influenza HA provided experimental evidence for “metastable” “native” protein structures to reconcile with the widely accepted thermodynamic views on protein folding beautifully illustrated experimentally by Anfinsen (1972). However, the fact that till date all experiments with biological membranes and compartments use detergents as a measure of “100%” disruptions show that membranes systems (both natural and model) are predominantly in a metastable state. Theoretically, post-disruption removal of detergent is not possible due to formation of newer equilibrium structures that would include detergent molecules. In this context, the following observations from literature are quite informative—
- a.
Bentz et al. (1983b) state It is important to mention that the equilibrium product for PS vesicles in a concentration of Ca2+ sufficient to induce fusion is a massive anhydrous structure called a cochleate ……. whose length is of the order of µm. Clearly, all of the vesicle contents will be leaked to the medium when the cochleates are eventually formed. However, the aggregation and fusion of two PS vesicles are nowhere near this equilibrium state, which is why the PS vesicle system (and other lipid mixtures such as PS/PC (phosphatidylcholine) …… can be used to study the fusion of two bilayers.
- b.
Lentz et al. (1987) state …. kinetic analysis of our data demonstrates, first, that small vesicle preparations should be used within a few hours of size fractionation. Second, the substantial increase in fusion rate with temperature (see Figure 6) indicates a large activation energy for the rate-limiting step of the process. From our data, we estimate this to be 30–40 kcal/mol. The large magnitude of this activation energy suggests that close juxtaposition of vesicle bilayers …… may be the rate-determining step for spontaneous fusion of SUV. Third ….. from a practical point of view, this means that when storage is necessary, SUV should be stored just above their phase transition. Finally, we note that only the smallest, most highly curved vesicles fused to form intermediate-sized vesicles. The intermediate-sized vesicles appear to be stable above their phase transition, although our data do not rule out the possibility that these, too, fuse, but at a much slower rate than do the highly curved species.
- a.
- (3)
Cells and biological systems in general do not operate at optimum temperatures (or pH) for all protein constituents. There are thermodynamic windows for operation of proteins in living systems, which are not optimized toward any single protein or function (Ghosh and Dill 2010). The same is applicable to biological membranes (in terms of phase transitions and properties of individual constituents).
- (4)
Consideration of only thermodynamic windows for operations in living systems is also highly misleading—kinetic windows for operation of proteins, whether from the perspectives of chemical operations of the central dogma or from the perspectives of diffusion, are equally important (Dill et al. 2011). The same is applicable for “functional” membrane assemblies that, while being thermodynamically unstable (in time scales of cellular operations), have kinetic windows for stabilization by “third-party” components.
The above highlights enough reasons to consider kinetic explorations of membrane assemblies and their constituents in biological membrane remodeling, rather than relying primarily on thermodynamic considerations. Thus, we end with Fig. 4 that couples possible kinetic transitions in membranes with thermodynamics. Based on existing biological membranes, it is clear that assemblies of amphipaths are “stabilized” by proteins (and other non-lipidic components) to form dynamic, yet (meta)stable, structures away from equilibrium configurations in aqueous systems. In fact, metastability of these structures is essential for operational dynamics of living systems—a very recent example of this comes from enzymatic hydrolysis of ectodomains of membrane proteins on avian erythrocytes revealing a novel non-montonic osmotic behavior of biological membranes that stayed undiscovered in spite of nearly 100 years of literature (Singh et al. 2019). To conclude, we propose that repeated de novo formation of metastable membrane structures (i.e., mimicking replication), with “third-party” constituents (due to their availability) were not only crucial for origins of life but also continue to offer morphological replication and/or functional mechanisms in modern life forms, independent of the central dogma. In fact, very recent experimental work exploring membrane assemblies and structures lends strong support to the above proposal (Steinkühler et al. 2020, 2021).
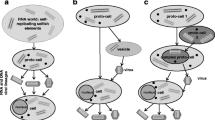
Theory on origins of life and biological replication independent of central dogma. A De novo appearance of a micelle and “replication” of the micelle based on law of mass action—a proposal on origins of life and evolution of biological systems from a world of biological membranes (see Mittal et al. 2020 for details). B Unilamellar compartments with bilayer formations from similar lipids, binary mixtures, and a heterogenous systems with lipids “third-party” components. C Free energy diagrams for hypothetical systems shown in A (red) and B (black). The arrows represent kinetic windows during thermodynamic transitions toward stabilization of membrane formations above the possible lowest energy states. The free energy curves also represent relative free energies of respective panels in A and B. The figure is drawn only for illustrative purposes and is not to any scale
References
Ada GL, Perry BT (1958) Properties of the nucleic acid of the Ryan strain of filamentous influenza virus. Microbiol 19(1):40–54. https://doi.org/10.1099/00221287-19-1-40
Adolf F, Rhiel M, Hessling B, Gao Q, Hellwig A, Béthune J, Wieland FT (2019) Proteomic profiling of mammalian COPII and COPI vesicles. Cell Rep 26(1):250-265.e5. https://doi.org/10.1016/j.celrep.2018.12.041
Aeffner S, Reusch T, Weinhausen B, Salditt T (2012) Energetics of stalk intermediates in membrane fusion are controlled by lipid composition. Proc Natl Acad Sci 109(25):E1609–E1618. https://doi.org/10.1073/pnas.1119442109
Agutter PS (2011) Stoichiometry-driven protein folding: a comment. J Biomol Struct Dyn 28(4):643–674. https://doi.org/10.1080/073911011010524974
Almeida PF (2014) The many faces of lipid rafts. Biophys J 106(9):1841–1843. https://doi.org/10.1016/j.bpj.2014.03.018
Alsaad KO, Hajeer AH, Al Balwi M, Al Moaiqel M, Al Oudah N, Al Ajlan A, AlJohani S, Alsolamy S, Gmati GE, Balkhy H, Al-Jahdali HH (2018) Histopathology of Middle East respiratory syndrome coronovirus (MERS-CoV) infection–clinicopathological and ultrastructural study. Histopathology 72(3):516–524. https://doi.org/10.1111/his.13379
Anfinsen CB (1972) Studies on the principles that govern folding of protein chains (Nobel Lecture, December 11, 1972). Nobelstiftelsen. Nobelprize.org: the Official Website of the Nobel Prize. Available at https://www.nobelprize.org/uploads/2018/06/anfinsen-lecture.pdf. Accessed 20 Jan 2022
Anilionis A, Wunner WH, Curtis PJ (1981) Structure of the glycoprotein gene in rabies virus. Nature 294(5838):275–278. https://doi.org/10.1038/294275a0
Antic D, Wright KE, Kang CY (1992) Maturation of Hantaan virus glycoproteins G1 and G2. Virology 189(1):324–328. https://doi.org/10.1016/0042-6822(92)90709-X
Backovic M, Jardetzky TS (2009) Class III viral membrane fusion proteins. Curr Opin Struct Biol 19(2):189–196. https://doi.org/10.1016/j.sbi.2009.02.012
Bangham AD (1972) Model membranes. Chem Phys Lipids 8:386–392. https://doi.org/10.1016/0009-3084(72)90069-2
Bangham AD, Horne RW (1964) Negative staining of phospholipids and their structural modification by surface-active agents as observed in the electron microscope. J Mol Biol 8(5):660–668. https://doi.org/10.1016/S0022-2836(64)80115-7
Bangham AD, Standish MM, Weissmann G (1965) The action of steroids and streptolysin S on the permeability of phospholipid structures to cations. J Mol Biol 13(1):253–259. https://doi.org/10.1016/S0022-2836(65)80094-8
Bansal S, Mittal A (2013) Extracting curvature preferences of lipids assembled in flat bilayers shows possible kinetic windows for genesis of bilayer asymmetry and domain formation in biological membranes. J Membr Biol 246(7):557–570. https://doi.org/10.1007/s00232-013-9568-1
Bansal S, Mittal A (2015) A statistical anomaly indicates symbiotic origins of eukaryotic membranes. Mol Biol Cell 26:1238–1248. https://doi.org/10.1091/mbc.E14-06-1078
Barlowe C, Orci L, Yeung T, Hosobuchi M, Hamamoto S et al (1994) COPII: a membrane coat formed by Sec proteins that drive vesicle budding from the endoplasmic reticulum. Cell 77:895–907. https://doi.org/10.1016/0092-8674(94)90138-4
Baumgart T, Hess ST, Webb WW (2003) Imaging coexisting fluid domains in biomembrane models coupling curvature and line tension. Nature 425(6960):821–824. https://doi.org/10.1038/nature02013
Beattie ME, Veatch SL, Stottrup BL, Keller SL (2005) Sterol structure determines miscibility versus melting transitions in lipid vesicles. Biophys J 89(3):1760–1768. https://doi.org/10.1529/biophysj.104.049635
Bentz J (2000a) Minimal aggregate size and minimal fusion unit for the first fusion pore of influenza hemagglutinin-mediated membrane fusion. Biophys J 78:227–245. https://doi.org/10.1016/S0006-3495(00)76587-8
Bentz J (2000b) Membrane fusion mediated by coiled coils: a hypothesis. Biophys J 78:886–900. https://doi.org/10.1016/S0006-3495(00)76646-X
Bentz J, Mittal A (2000) Deployment of membrane fusion protein domains during fusion. Cell Biol Int 24:819–838. https://doi.org/10.1006/cbir.2000.0632
Bentz J, Mittal A (2003) Architecture of the influenza hemagglutinin membrane fusion site. Biochim Biophys Acta 1614:24–35. https://doi.org/10.1016/S0005-2736(03)00160-3
Bentz J, Nir S (1981a) Aggregation of colloidal particles modeled as a dynamical process. Proc Natl Acad Sci USA 78:1634-1637. https://doi.org/10.1073/pnas.78.3.1634
Bentz J, Nir S (1981b) Mass action kinetics and equilibria of reversible aggregation. J Chem Soc Faraday Trans I 77:1249–1275. https://doi.org/10.1039/F19817701249
Bentz J, Düzgüneş N, Nir S (1983a) Kinetics of divalent cation induced fusion of phosphatidylserine vesicles: correlation between fusogenic capacities and binding affinities. Biochemistry 22(14):3320–3330
Bentz J, Nir S, Wilschut J (1983b) Mass-action kinetics of vesicle aggregation and fusion. Colloids Surf 6(4):333–363. https://doi.org/10.1016/0166-6622(83)80026-2
Bentz J, Ellens H, Alford D (1990) An architecture for the fusion site of Influenza hemagglutinin. FEBS Lett 276(1–2):1–5. https://doi.org/10.1016/0014-5793(90)80492-2
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The protein data bank. Nucleic Acids Res 28:235–242. https://doi.org/10.1093/nar/28.1.235
Bhatia T, Husen P, Brewer J, Bagatolli LA, Hansen PL, Ipsen JH, Mouritsen OG (2015) Preparing giant unilamellar vesicles (GUVs) of complex lipid mixtures on demand: mixing small unilamellar vesicles of compositionally heterogeneous mixtures. Biochim Biophys Acta 1848(12):3175–80. https://doi.org/10.1016/j.bbamem.2015.09.020
Bhatia T, Agudo-Canalejo J, Dimova R, Lipowsky R (2018) Membrane nanotubes increase the robustness of giant vesicles. ACS Nano 12(5):4478–4485. https://doi.org/10.1021/acsnano.8b00640
Bhatia T, Christ S, Steinkühler J, Dimova R, Lipowsky R (2020) Simple sugars shape giant vesicles into multispheres with many membrane necks. Soft Matter 16(5):1246–1258. https://doi.org/10.1039/c9sm01890e
Biswas S, Yin SR, Blank PS, Zimmerberg J (2008) Cholesterol promotes hemifusion and pore widening in membrane fusion induced by influenza hemagglutinin. J Gen Physiol 131(5):503–513. https://doi.org/10.1085/jgp.200709932
Blott EJ, Griffiths GM (2002) Secretory lysosomes. Nat Rev Mol Cell Biol 3(2):122–131. https://doi.org/10.1038/nrm732
Blumenthal R, Sarkar DP, Durell S, Howard DE, Morris SJ (1996) Dilation of the influenza hemagglutinin fusion pore revealed by the kinetics of individual fusion events. J Cell Biol 135:63–71. https://doi.org/10.1083/jcb.135.1.63
Borza CM, Hutt-Fletcher LM (2002) Alternate replication in B cells and epithelial cells switches tropism of Epstein-Barr virus. Nat Med 8(6):594–599. https://doi.org/10.1038/nm0602-594
Bosch BJ, Van der Zee R, De Haan CA, Rottier PJ (2003) The coronavirus spike protein is a class I virus fusion protein: structural and functional characterization of the fusion core complex. J Virol 77(16):8801–8811. https://doi.org/10.1128/JVI.77.16.8801-8811.2003
Brunner Y, Schvartz D, Couté Y, Sanchez JC (2009) Proteomics of regulated secretory organelles. Mass Spectrom Rev 28:844–867. https://doi.org/10.1002/MAS.20211
Bullough PA, Hughson FM, Skehel JJ, Wiley DC (1994) Structure of Influenza Hameagglutinin at the pH of membrane fusion. Nature 371:37–43. https://doi.org/10.1038/371037a0
Calder LJ, Wasilewski S, Berriman JA, Rosenthal PB (2010) Structural organization of a filamentous influenza A virus. Proc Nat Acad Sci 107(23):10685–10690. https://doi.org/10.1073/pnas.1002123107
Carr CM, Kim PS (1993) A spring-loaded mechanism for the conformational change of influenza hemagglutinin. Cell 73(4):823–832. https://doi.org/10.1016/0092-8674(93)90260-w
Casares D, Escribá PV, Rosselló CA (2019) Membrane lipid composition: Effect on membrane and organelle structure, function and compartmentalization and therapeutic avenues. Int J Mol Sci 20(9):2167. https://doi.org/10.3390/ijms20092167
Cech TR (2012) The RNA worlds in context. Cold Spring Harb Perspect Biol 4(7):a006742. https://doi.org/10.1101/cshperspect.a006742
Chambers TJ, Hahn CS, Galler R, Rice CM (1990) Flavivirus genome organization, expression, and replication. Annu Rev Microbiol 44(1):649–688. https://doi.org/10.1146/annurev.mi.44.100190.003245
Chan DC, Fass D, Berger JM, Kim PS (1997) Core structure of gp41 from the HIV envelope glycoprotein. Cell 89(2):263–273. https://doi.org/10.1016/S0092-8674(00)80205-6
Chargaff E (1950) Chemical specificity of nucleic acids and mechanism of their enzymatic degradation. Experientia 6:201–209. https://doi.org/10.1007/BF02173653
Chen Y, Maguire T, Marks RM (1996) Demonstration of binding of dengue virus envelope protein to target cells. J Virol 70(12):8765–8772. https://doi.org/10.1128/jvi.70.12.8765-8772.1996
Chernomordik LV, Melikyan GB, Chizmadzhev YA (1987) Biomembrane fusion: a new concept derived from model studies using two interacting planar lipid bilayers. Biochim Biophys Acta 906(3):309–352. https://doi.org/10.1016/0304-4157(87)90016-5
Chernomordik LV, Leikina E, Frolov V, Bronk P, Zimmerberg J (1997) An early stage of membrane fusion mediated by the low pH conformation of influenza hemagglutinin depends upon membrane lipids. J Cell Biol 136:81–93. https://doi.org/10.1083/jcb.136.1.81
Chernomordik LV, Frolov VA, Leikina E, Bronk P, Zimmerberg J (1998) The pathway of membrane fusion catalyzed by influenza hemagglutinin: restriction of lipids, hemifusion, and lipid fusion pore formation. J Cell Biol 140:1369–1382. https://doi.org/10.1083/jcb.140.6.1369
Choi HK, Tong L, Minor W, Dumas P, Boege U, Rossmann MG, Wengler G (1991) Structure of Sindbis virus core protein reveals a chymotrypsin-like serine proteinase and the organization of the virion. Nature 354(6348):37–43. https://doi.org/10.1038/354037a0
Cocucci E, Meldolesi J (2011) Ectosomes Current Biol 21(23):R940–R941. https://doi.org/10.1016/j.cub.2011.10.011
Cocucci E, Meldolesi J (2015) Ectosomes and exosomes: shedding the confusion between extracellular vesicles. Trends Cell Biol 25(6):364–372. https://doi.org/10.1016/j.tcb.2015.01.004
Cohen JS, Jardetzky O, Epstein H et al (1981) Phospholipid vesicle formation and transmembrane protein incorporation using octyl glucoside. Biochemistry 20:833–840. https://doi.org/10.1021/BI00507A028
Colf LA, Juo ZS, Garcia KC (2007) Structure of the measles virus hemagglutinin. Nat Struct Mol Biol 14(12):1227–1228. https://doi.org/10.1038/nsmb1342
Dadonaite B, Vijayakrishnan S, Fodor E, Bhella D, Hutchinson EC (2016) Filamentous influenza viruses. J Gen Virol 97(8):1755. https://doi.org/10.1099/jgv.0.000535
Dalrymple JM, Schlesinger S, Russell PK (1976) Antigenic characterization of two Sindbis envelope glycoproteins separated by isoelectric focusing. Virology 69(1):93–103. https://doi.org/10.1016/0042-6822(76)90197-5
Danchin A (2014) The emergence of the first cells. Rev Cell Biol Mol Suppl. https://doi.org/10.1002/3527600906.mcb.20130025
Danieli T, Pelletier SL, Henis YI, White JM (1996) Membrane fusion mediated by the influenza virus hemagglutinin requires the concerted action of at least three hemagglutinin trimers. J Cell Biol 133:559–569. https://doi.org/10.1083/jcb.133.3.559
Di Vizio D, Kim J, Hager MH, Morello M, Yang W, Lafargue CJ, True LD, Rubin MA, Adam RM, Beroukhim R, Demichelis F, Freeman MR (2009) Oncosome formation in prostate cancer: association with a region of frequent chromosomal deletion in metastatic disease. Cancer Res 69(13):5601–5609. https://doi.org/10.1158/0008-5472.CAN-08-3860
Dill KA, Ghosh K, Schmit JD (2011) Physical limits of cells and proteomes. Proc Natl Acad Sci USA 108(44):17876–17882. https://doi.org/10.1073/pnas.1114477108
Dixon LK, Chapman D (2008) African Swine fever virus. In: Mahy BWJ, Van Regenmortel MHV (eds) Encyclopedia of virology, 3rd edn. Academic Press, Cambridge, pp 43–51
Doxsey SJ, Sambrook J, Helenius A, White J (1985) An efficient method for introducing macromolecules into living cells. J Cell Biol 101(1):19–27. https://doi.org/10.1083/jcb.101.1.19
Elford WJ, Andrewes CH, Tang FF (1936) The sizes of the viruses of human and swine influenza, as determined by ultra-filtration. Br J Exp Pathol 17(1):51–53
Elizondo E, Larsen J, Hatzakis NS et al (2012) Influence of the preparation route on the supramolecular organization of lipids in a vesicular system. J Am Chem Soc 134:1918–1921. https://doi.org/10.1021/ja2086678
Ellens H, Siegel DP, Alford D et al (1989) Membrane fusion and inverted phases. Biochemistry 28:3692–3703. https://doi.org/10.1021/BI00435A011
Ellens H, Bentz J, Mason D, Zhang F, White JM (1990) Fusion of influenza hemagglutinin-expressing fibroblasts with glycophorin bearing liposomes: role of hemagglutinin surface density. Biochemistry 29:9697–9707. https://doi.org/10.1021/bi00493a027
Feldman SA, Hendry RM, Beeler JA (1999) Identification of a linear heparin binding domain for human respiratory syncytial virus attachment glycoprotein G. J Virol 73(8):6610–6617. https://doi.org/10.1128/jvi.73.8.6610-6617.1999
Feldmann H, Jones S, Klenk HD, Schnittler HJ (2003) Ebola virus: from discovery to vaccine. Nat Rev Immunol 3(8):677–685. https://doi.org/10.1038/nri1154
Forrester AU, Farrell H, Wilkinson GA, Kaye JA, Davis-Poynter NI, Minson TO (1992) Construction and properties of a mutant of herpes simplex virus type 1 with glycoprotein H coding sequences deleted. J Virol 66(1):341–348. https://doi.org/10.1128/jvi.66.1.341-348.1992
Füzik T, Formanová P, Růžek D, Yoshii K, Niedrig M, Plevka P (2018) Structure of tick-borne encephalitis virus and its neutralization by a monoclonal antibody. Nat Commun 9(1):1–1. https://doi.org/10.1038/s41467-018-02882-0
Ganar K, Das M, Sinha S, Kumar S (2014) Newcastle disease virus: current status and our understanding. Virus Res 184:71–81. https://doi.org/10.1016/j.virusres.2014.02.016
Ganley IG, Carroll K, Bittova L, Pfeffer S (2004) Rab9 GTPase regulates late endosome size and requires effector interaction for its stability. Mol Biol Cell 15:5420–5430. https://doi.org/10.1091/MBC.E04-08-0747
Garoff H, Simons K, Renkonen O (1974) Isolation and characterization of the membrane proteins of Semliki Forest virus. Virology 61(2):493–504. https://doi.org/10.1016/0042-6822(74)90285-2
Gattegno T, Mittal A, Valansi C, Nguyen KC, Hall DH, Chernomordik LV, Podbilewicz B (2007) Genetic control of fusion pore expansion in the epidermis of Caenorhabditis elegans. Mol Biol Cell 8(4):1153–1166. https://doi.org/10.1091/mbc.e06-09-0855
Gaudin Y, Tuffereau C, Segretain D, Knossow M, Flamand A (1991) Reversible conformational changes and fusion activity of rabies virus glycoprotein. J Virol 65(9):4853–4859. https://doi.org/10.1128/JVI.65.9.4853-4859.1991
Ge P, Tsao J, Schein S, Green TJ, Luo M, Zhou ZH (2010) Cryo-EM model of the bullet-shaped vesicular stomatitis virus. Science 327(5966):689–693. https://doi.org/10.1126/science.1181766
Gething MJ, Doms RW, York D, White J (1986) Studies on the mechanism of membrane fusion: site-specific mutagenesis of the hemagglutinin of influenza virus. J Cell Biol 102:11–23. https://doi.org/10.1083/jcb.102.1.11
Ghosh K, Dill K (2010) Cellular proteomes have broad distributions of protein stability. Biophys J 99(12):3996–4002. https://doi.org/10.1016/j.bpj.2010.10.036
Gondré-Lewis MC, Park JJ, Loh YP (2012) Cellular mechanisms for the biogenesis and transport of synaptic and dense-core vesicles. Int Rev Cell and Mol Biol 299:27–115. https://doi.org/10.1016/B978-0-12-394310-1.00002-3
Greco V, Hannus M, Eaton S (2001) Argosomes: a potential vehicle for the spread of morphogens through epithelia. Cell 106(5):633–645. https://doi.org/10.1016/S0092-8674(01)00484-6
Griffith F (1928) The significance of pneumococcal types. J Hygiene 27(2):113–159. https://doi.org/10.1017/S0022172400031879
Griffiths G (2002) What’s special about secretory lysosomes? Sem Cell Dev Biol 13(4):279–284. https://doi.org/10.1016/S1084-9521(02)00057-5
Grünewald K, Desai P, Winkler DC, Heymann JB, Belnap DM, Baumeister W, Steven AC (2003) Three-dimensional structure of herpes simplex virus from cryo-electron tomography. Science 302(5649):1396–1398. https://doi.org/10.1126/science.1090284
Guerrier-Takada C, Gardiner K, Marsh T, Pace N, Altman S (1983) The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell 35:849–857. https://doi.org/10.1016/0092-8674(83)90117-4
Günther S, Lenz O (2004) Lassa virus. Crit Rev Clin Lab Sci 41:339–390. https://doi.org/10.1080/10408360490497456
Gupta V, Gupta R, Grover R, Khanna R, Jangra V, Mittal A (2008) Delivery of molecules to cancer cells using liposomes from bacterial cultures. J Nanosci Nanotechnol 8:2328–2333. https://doi.org/10.1166/jnn.2008.18273
Harris A, Cardone G, Winkler DC, Heymann JB, Brecher M, White JM, Steven AC (2006) Influenza virus pleiomorphy characterized by cryoelectron tomography. Proc Nat Acad Sci 103(50):19123–19127. https://doi.org/10.1073/pnas.0607614103
Harvey RA, Champe PC, Fisher BD, Strohl WA (2007) Lippincott’s illustrated reviews: Microbiology. Lippincott Williams & Wilkins, Philadelphia
Hata Y, Slaughter CA, Sudhof TC (1993) Synaptic vesicle fusion complex contains unc-18 homologue bound to syntaxin. Nature 366:347–351. https://doi.org/10.1038/366347a0
Hauser P, Wang S, Didenko VV (2017) Apoptotic bodies: selective detection in extracellular vesicles. In: Kalyuzhny A (ed) Signal transduction immunohistochemistry. Methods in molecular biology, vol 1554. Humana Press, New York, pp 193–200
Haywood AM (1974) Fusion of Sendai viruses with model membranes. J Mol Biol 87:625–628. https://doi.org/10.1016/0022-2836(74)90107-7
Helenius A, Kartenbeck J, Simons K, Fries E (1980) On the entry of Semliki Forest virus into BHK-21 cells. J Cell Biol 84:404–420. https://doi.org/10.1083/jcb.84.2.404
Herold BC, WuDUNN DA, Soltys N, Spear PG (1991) Glycoprotein C of herpes simplex virus type 1 plays a principal role in the adsorption of virus to cells and in infectivity. J Virol 65(3):1090–1098. https://doi.org/10.1128/jvi.65.3.1090-1098.1991
Herrmann A, Clague MJ, Blumenthal R (1993) Role of target membrane structure in fusion with influenza virus: effect of modulating erythrocyte transbilayer phospholipid distribution. Membr Biochem 10(1):3–15. https://doi.org/10.1083/jcb.84.2.404
Hershey AD, Chase M (1952) Independent functions of viral protein and nucleic acid in growth of bacteriophage. J Gen Physiol 36(1):39–56. https://doi.org/10.1085/jgp.36.1.39
Hess ST, Kumar M, Verma A, Farrington J, Kenworthy A, Zimmerberg J (2005) Quantitative electron microscopy and fluorescence spectroscopy of the membrane distribution of influenza hemagglutinin. J Cell Biol 169(6):965–976. https://doi.org/10.1083/jcb.200412058
Hess ST, Gould TJ, Gudheti MV, Maas SA, Mills KD, Zimmerberg J (2007) Dynamic clustered distribution of hemagglutinin resolved at 40 nm in living cell membranes discriminates between raft theories. Proc Natl Acad Sci USA 104(44):17370–17375. https://doi.org/10.1073/pnas.0708066104
Hoekstra D, de Boer T, Klappe K, Wilschut J (1984) Fluorescence method for measuring the kinetics of fusion between biological membranes. Biochemistry 23(24):5675–5681. https://doi.org/10.1021/bi00319a002
Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S, Schiergens TS, Herrler G, Wu NH, Nitsche A, Müller MA (2020) SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181(2):271–280. https://doi.org/10.1016/j.cell.2020.02.052
Holt OJ, Gallo F, Griffiths GM (2006) Regulating secretory lysosomes. J Biochem 140(1):7–12. https://doi.org/10.1093/jb/mvj126
Howe C, Morgan C, Cyr CD, Hsu KC, Rose H (1967) Morphogenesis of type 2 parainfluenza virus examined by light and electron microscopy. J Virol 1(1):215–237. https://doi.org/10.1128/jvi.1.1.215-237.1967
Hu XL, Ray R, Compans RW (1992) Functional interactions between the fusion protein and hemagglutinin-neuraminidase of human parainfluenza viruses. J Virol 66(3):1528–1534. https://doi.org/10.1128/jvi.66.3.1528-1534.1992
Huotari J, Helenius A (2011) Endosome maturation. EMBO J 30:3481–3500. https://doi.org/10.1038/EMBOJ.2011.286
Israelachvili JN, Mitchell DJ, Ninham BW (1976) Theory of self assembly of hydrocarbon amphiphiles into micelles and bilayers. J Chem Soc Faraday Trans 72:1525–1568. https://doi.org/10.1039/F29767201525
Jacobson K, Sheets ED, Simson R (1995) Revisiting the fluid mosaic model of membranes. Science 268(5216):1441–1442. https://doi.org/10.1126/science.7770769
Jaiswal R, Sedger LM (2019) Intercellular vesicular transfer by exosomes, microparticles and oncosomes—implications for cancer biology and treatments. Front Oncol 9:125. https://doi.org/10.3389/fonc.2019.00125
Jaquinod M, Villiers F, Kieffer-Jaquinod S, Hugouvieux V, Bruley C, Garin J, Bourguignon J (2007) A proteomics dissection of Arabidopsis thaliana vacuoles isolated from cell culture. Mol Cell Proteomics 6(3):394–412. https://doi.org/10.1074/mcp.M600250-MCP200
Johnson L, Gupta AK, Ghafoor A, Akin D, Bashir R (2006) Characterization of vaccinia virus particles using microscale silicon cantilever resonators and atomic force microscopy. Sens Actuators B Chem 115(1):189–197. https://doi.org/10.1016/j.snb.2005.08.047
Jovic M, Sharma M, Rahajeng J, Caplan S (2010) The early endosome: a busy sorting station for proteins at the crossroads. Histol Histopathol 25(1):99. https://doi.org/10.14670/hh-25.99
Kalra H, Simpson RJ, Ji H, Aikawa E, Altevogt P, Askenase P, Bond VC, Borras FE, Breakefield X, Budnik V, Buzas E, Camussi G, Clayton A, Cocucci E, Falcon-Perez JM, Gabrielsson S, Gho YS, Gupta D, Harsha HC, Hendrix A, Hill AF, Inaal JM, Jenster G, Kiang LS, Kramer-Albers E-M, Llorente A, Lotvall J, Mincheva-Nilsson L, Nazarenko I, Nieuwland R, Nolte-’t Hoen ENM, Pandey A, Patel T, Piper MG, Pluchino S, Prasad TSK, Rajendran L, Raposo G, Record M, Reid GE, Sanchez-Madrid F, Schiffelers RM, Siljander P, Stoorvogel W, Taylor D, Thery C, Valadi H, van Balkom BWM, Vazquez J, Vidal M, Yanez-Mo M, Zoeller M, Mathivanan S (2012) Vesiclepedia: a compendium for extracellular vesicles with continuous community annotation. PLoS Biol 12:e1001450. https://doi.org/10.1371/journal.pbio.1001450
Kamal MA, Raghunathan VA (2012) Phase behavior of phospholipid–phytosterol membranes. Soft Matter 8:8952–8958. https://doi.org/10.1039/C2SM25912E
Kanai R, Kar K, Anthony K, Gould LH, Ledizet M, Fikrig E, Marasco WA, Koski RA, Modis Y (2006) Crystal structure of West Nile virus envelope glycoprotein reveals viral surface epitopes. J Virol 80(22):11000–11008. https://doi.org/10.1128/JVI.01735-06
Kanaseki T, Kadota K (1969) A morphological study of the coated vesicle isolated from the nerve endings of the guinea pig brain, with special reference to the mechanism of membrane movements. J Cell Biol 42(1):202–220. https://doi.org/10.1083/jcb.42.1.202
Karatekin E, Sandre O, Guitouni H, Borghi N, Puech PH, Brochard-Wyart F (2003) Cascades of transient pores in giant vesicles: line tension and transport. Biophys J 84(3):1734–1749. https://doi.org/10.1016/S0006-3495(03)74981-9
Karlsson M, Lundin S, Dahlgren U, Kahu H, Pettersson I, Telemo E (2001) “Tolerosomes” are produced by intestinal epithelial cells. Eur J Immunol 31(10):2892–2900. https://doi.org/10.1002/1521-4141(2001010)31:10%3c2892::AID-IMMU2892%3e3.0.CO;2-I
Karmakar S, Raghunathan VA (2005) Structure of phospholipid-cholesterol membranes: an x-ray diffraction study. Phys Rev E Stat Nonlin Soft Matter Phys 71:061924. https://doi.org/10.1103/PhysRevE.71.061924
Karunakaran S, Fratti RA (2013) The lipid composition and physical properties of the yeast vacuole affect the hemifusion–fusion transition. Traffic 14(6):650–662. https://doi.org/10.1111/tra.12064
Keerthikumar S, Chisanga D, Ariyaratne D, Al Saffar H, Anand S, Zhao K, Samuel M, Pathan M, Jois M, Chilamkurti N, Gangoda L, Mathivanan S (2016) ExoCarta: a web-based compendium of exosomal cargo. J Mol Biol 428(4):688–692. https://doi.org/10.1016/j.jmb.2015.09.019
Kelley JM, Emerson SU, Wagner RR (1972) The glycoprotein of vesicular stomatitis virus is the antigen that gives rise to and reacts with neutralizing antibody. J Virol 10(6):1231–1235. https://doi.org/10.1128/jvi.10.6.1231-1235.1972
Kim T, Gondré-Lewis MC, Arnaoutova I, Loh YP (2006) Dense-core secretory granule biogenesis. Physiology 21:124–133. https://doi.org/10.1152/physiol.00043.2005
Kostyuchenko VA, Lim EX, Zhang S, Fibriansah G, Ng TS, Ooi JS, Shi J, Lok SM (2016) Structure of the thermally stable Zika virus. Nature 533(7603):425–428. https://doi.org/10.1038/nature17994
Kozlov MM, Markin VS (1983) Vozmozhnyĭ mekhanizm sliiania membran [Possible mechanism of membrane fusion]. Biofizika. 28(2):242–247
Kruger K, Grabowski PJ, Zaug AJ, Sands J, Gottschling DE, Cech TR (1982) Self-splicing RNA: autoexcision and autocyclization of the ribosomal RNA intervening sequence of tetrahymena. Cell 31:147–157. https://doi.org/10.1016/0092-8674(82)90414-7
Kruger S, Abd Elmageed ZY, Hawke DH, Wörner PM, Jansen DA, Abdel-Mageed AB, Alt EU, Izadpanah R (2014) Molecular characterization of exosome-like vesicles from breast cancer cells. BMC Cancer. https://doi.org/10.1186/1471-2407-14-44
Kuhn JH, Radoshitzky SR, Guth AC, Warfield KL, Li W, Vincent MJ, Towner JS, Nichol ST, Bavari S, Choe H, Aman MJ (2006) Conserved receptor-binding domains of Lake Victoria marburgvirus and Zaire ebolavirus bind a common receptor. J Biol Chem 281(23):15951–15958. https://doi.org/10.1074/jbc.M601796200
Kühne W (1877) Über das Verhalten verschiedener organisirter und sog. ungeformter Fermente. Verhandlungen des Naturhistorisch-medicinischen Vereins zu Heidelberg 1(3):190–193
Kuznetsov IA, Kuznetsov AV (2017) How dense core vesicles are delivered to axon terminals–a review of modeling approaches. Model Microscale Transp Biol Process. https://doi.org/10.1016/B978-0-12-804595-4.00013-4
Kuznetsov YG, Xiao C, Sun S, Raoult D, Rossmann M, McPherson A (2010) Atomic force microscopy investigation of the giant mimivirus. Virology 404(1):127–137. https://doi.org/10.1016/j.virol.2010.05.007
Landolfo S, Gariglio M, Gribaudo G, Lembo D (2003) The human cytomegalovirus. Pharmacol Ther 98(3):269–297. https://doi.org/10.1016/S0163-7258(03)00034-2
Lawrence MC, Borg NA, Streltsov VA, Pilling PA, Epa VC, Varghese JN, McKimm-Breschkin JL, Colman PM (2004) Structure of the haemagglutinin-neuraminidase from human parainfluenza virus type III. J Mol Biol 335(5):1343–1357. https://doi.org/10.1016/j.jmb.2003.11.032
Lee J, Kreutzberger AJB, Odongo L, Nelson EA, Nyenhuis DA, Kiessling V, Liang B, Cafiso DS, White JM, Tamm LK (2021) Ebola virus glycoprotein interacts with cholesterol to enhance membrane fusion and cell entry. Nat Struct Mol Biol 28(2):181–189. https://doi.org/10.1038/s41594-020-00548-4
Lei G, MacDonald RC (2003) Lipid bilayer vesicle fusion: intermediates captured by high-speed microfluorescence spectroscopy. Biophys J 85(3):1585–1599. https://doi.org/10.1016/S0006-3495(03)74590-1
Leikina E, Mittal A, Cho M, Melikov K, Kozlov MM, Chernomordik LV (2004) Influenza hemagglutinins outside of the contact zone are necessary for fusion pore expansion. J Biol Chem 279:26526–26532. https://doi.org/10.1074/jbc.M401883200
Lentz BR, Carpenter TJ, Alford DR (1987) Spontaneous fusion of phosphatidylcholine small unilamellar vesicles in the fluid phase. Biochemistry 26:5389–5397. https://doi.org/10.1021/bi00391a026
Ligas MW, Johnson DC (1988) A herpes simplex virus mutant in which glycoprotein D sequences are replaced by beta-galactosidase sequences binds to but is unable to penetrate into cells. J Virol 62(5):1486–1494. https://doi.org/10.1128/jvi.62.5.1486-1494.1988
Lübke T, Lobel P, Sleat DE (2009) Proteomics of the lysosome. Biochim Biophys Acta BBA Mol Cell Res 1793(4):625–35. https://doi.org/10.1016/j.bbamcr.2008.09.018
Malhotra V, Orci L, Glick BS, Block MR, Rothman JE (1988) Role of an N-ethylmaleimide-sensitive transport component in promoting fusion of transport vesicles with cisternae of the Golgi stack. Cell 54:221–227. https://doi.org/10.1016/0092-8674(88)90554-5
Malhotra V, Serafini T, Orci L, Shepard JS, Rothman JE (1989) Purification of a novel class of coated vesicles mediating biosynthetic protein-transport through the Golgi stack. Cell 58:329–336. https://doi.org/10.1016/0092-8674(89)90847-7
Mallat Z, Hugel B, Ohan J, Leseche G, Freyssinet JM, Tedgui A (1999) Shed membrane microparticles with procoagulant potential in human atherosclerotic plaques: a role for apoptosis in plaque thrombogenicity. Circulation 99:348–353. https://doi.org/10.1161/01.CIR.99.3.348
Mancini EJ, Clarke M, Gowen BE, Rutten T, Fuller SD (2000) Cryo-electron microscopy reveals the functional organization of an enveloped virus. Semliki Forest Virus Mol Cell 5(2):255–266. https://doi.org/10.1016/S1097-2765(00)80421-9
Marchiani S, Tamburrino L, Maoggi A, Vannelli GB, Forti G, Baldi E, Muratori M (2007) Characterization of M540 bodies in human semen: evidence that they are apoptotic bodies. Mol Hum Reprod 13(9):621–631. https://doi.org/10.1093/molehr/gam046
Mathivanan S, Fahner CJ, Reid GE, Simpson RJ (2012) ExoCarta 2012: database of exosomal proteins. RNA Lipids Nucleic Acids Res 40(D1):D1241–D1244. https://doi.org/10.1093/nar/gkr828
Matile P (1978) Biochemistry and function of vacuoles. Annu Rev Plant Physiol 29:193–213. https://doi.org/10.1146/ANNUREV.PP.29.060178.001205
Maurer SE, Deamer DW, Boncella JM, Monnard PA (2009) Chemical evolution of amphiphiles: glycerol monoacyl derivatives stabilize plausible prebiotic membranes. Astrobiology 9:979–987. https://doi.org/10.1089/AST.2009.0384
Meehan B, Rak J, di Vizio D (2016) Oncosomes—large and small: what are they, where they came from? J Extracell Vesicles 5(1):33109. https://doi.org/10.3402/JEV.V5.33109
Meers PR (2022) Membrane organization strategies in vesicular antibiotic delivery. J Membr Biol. https://doi.org/10.1007/s00232-021-00210-0
Melikyan GB, Niles W, Cohen FS (1995) The fusion kinetics of influenza hemagglutinin expressing cells to planar bilayer membranes is affected by HA surface density and host cell surface. J Gen Physiol 106:783–802. https://doi.org/10.1085/jgp.106.5.783
Mittal A (2009) Book Review: Protocells: bridging nonliving and living matter by Rasmussen et al. (eds). Curr Sci 96(11):1530–1532
Mittal A (2012) Self-generated and reproducible dynamics in “gene years” represent life. J Biomol Struct Dyn 29(4):609–611. https://doi.org/10.1080/073911012010525002
Mittal A, Acharya C (2012) Extracting signatures of spatial organization for biomolecular nanostructures. J Nanosci Nanotechnol 12(11):8249–8257. https://doi.org/10.1166/jnn.2012.6732
Mittal A, Acharya C (2013) Protein folding: Is it simply surface to volume minimization? J Biomol Struct Dyn 31:953–955. https://doi.org/10.1080/07391102.2012.748526
Mittal A, Bentz J (2001) Comprehensive kinetic analysis of influenza hemagglutinin-mediated membrane fusion: role of sialate binding. Biophys J 81:1521–1535. https://doi.org/10.1016/S0006-3495(01)75806-7
Mittal A, Grover R (2010) Self-assembly of biological membranes into 200–400 nm aqueous compartments. J Nanosci Nanotechnol 10:3085–3090. https://doi.org/10.1166/jnn.2010.2514
Mittal A, Jayaram B (2011a) Backbones of folded proteins reveal novel invariant amino acid neighborhoods. J Biomol Struct Dyn 28(4):443–454. https://doi.org/10.1080/073911011010524954
Mittal A, Jayaram B (2011b) The newest view on protein folding: stoichiometric and spatial unity in structural and functional diversity. J Biomol Struct Dyn 28(4):669–674. https://doi.org/10.1080/073911011010524984
Mittal A, Singh S (2018) Insights into eukaryotic evolution from transmembrane domain lengths. J Biomol Struct Dyn 36(8):2194–2200. https://doi.org/10.1080/07391102.2017.1345699
Mittal A, Leikina E, Bentz J, Chernomordik LV (2002a) Kinetics of influenza hemagglutinin-mediated membrane fusion as a function of technique. Anal Biochem 303:145–152. https://doi.org/10.1006/abio.2002.5590
Mittal A, Shangguan T, Bentz J (2002b) Measuring pKa of activation and pKi of inactivation for influenza hemagglutinin from kinetics of membrane fusion of virions and of HA expressing cells. Biophys J 83:2652–2666. https://doi.org/10.1016/S0006-3495(02)75275-2
Mittal A, Leikina E, Chernomordik LV, Bentz J (2003) Kinetically differentiating influenza hemagglutinin fusion and hemifusion machines. Biophys J 85:1713–1724. https://doi.org/10.1016/S0006-3495(03)74601-3
Mittal A, Jayaram B, Shenoy S, Bawa TS (2010) A stoichiometry driven universal spatial organization of backbones of folded proteins: are there Chargaff’s rules for protein folding? J Biomol Struct Dyn 28(2):133–142. https://doi.org/10.1080/07391102.2010.10507349
Mittal A, Bansal S, Changani AM (2020) On the origin of life and evolution of living systems from a world of biological membranes. In: Pontarotti P (ed) Evolutionary biology—a transdisciplinary approach. Springer, Cham. https://doi.org/10.1007/978-3-030-57246-4_8
Mittal A, Changani AM, Taparia S (2021a) What limits the primary sequence space of natural proteins? J Biomol Struct Dyn 38(15):4579–4583. https://doi.org/10.1080/07391102.2019.1682051
Mittal A, Changani AM, Taparia S (2021b) Unique and exclusive peptide signatures directly identify intrinsically disordered proteins from sequences without structural information. J Biomol Struct Dyn 39(8):2885–2893. https://doi.org/10.1080/07391102.2020.1756410
Mittal A, Changani AM, Taparia S, Goel D, Parihar A, Singh I (2021c) Structural disorder originates beyond narrow stoichiometric margins of amino acids in naturally occurring folded proteins. J Biomol Struct Dyn 39(7):2364–2375. https://doi.org/10.1080/07391102.2020.1751299
Modis Y (2014) Relating structure to evolution in class II viral membrane fusion proteins. Curr Opin Virol 5:34–41. https://doi.org/10.1016/j.coviro.2014.01.009
Morgan C, Howe C (1968) Structure and development of viruses as observed in the electron microscope. IX. Entry of parainfluenza I (Sendai) virus. J Virol 2:1122–1132. https://doi.org/10.1128/jvi.2.10.1122-1132.1968
Morris SJ, Sarkar DP, White JM, Blumenthal R (1989) Kinetics of pH-dependent fusion between 3T3 fibroblasts expressing influenza hemagglutinin and red blood cells: measurement by dequenching of fluorescence. J Biol Chem 264(7):3972–3978. https://doi.org/10.1016/S0021-9258(19)84948-7
Moscho A, Orwar O, Chiu DT, Modi BP, Zare RN (1996) Rapid preparation of giant unilamellar vesicles. Proc Natl Acad Sci USA 93(21):11443–11447. https://doi.org/10.1073/pnas.93.21.11443
Mouritsen OG, Bloom M (1984) Mattress model of lipid-protein interactions in membranes. Biophys J 46(2):141–153. https://doi.org/10.1016/S0006-3495(84)84007-2
Mufson MA, Örvell C, Rafnar B, Norrby E (1985) Two distinct subtypes of human respiratory syncytial virus. J Gen Virol 66(10):2111–2124. https://doi.org/10.1099/0022-1317-66-10-2111
Mukhopadhyay S, Kim BS, Chipman PR, Rossmann MG, Kuhn RJ (2003) Structure of west Nile virus. Science 302(5643):248. https://doi.org/10.1126/SCIENCE.1089316
Nagai Y, Klenk HD, Rott R (1976) Proteolytic cleavage of the viral glycoproteins and its significance for the virulence of Newcastle disease virus. Virology 72(2):494–508. https://doi.org/10.1016/0042-6822(76)90178-1
Nagle JF (2017) Experimentally determined tilt and bending moduli of single-component lipid bilayers. Chem Phys Lipids 205:18–24. https://doi.org/10.1016/j.chemphyslip.2017.04.006
Nanbo A, Noda T, Ohba Y (2018) Epstein-Barr virus acquires its final envelope on intracellular compartments with golgi markers. Front Microbiol 9:454. https://doi.org/10.3389/fmicb.2018.00454
Näslund TI, Gehrmann U, Qazi KR, Karlsson MCI, Gabrielsson S (2013) Dendritic cell-derived exosomes need to activate both T and B cells to induce antitumor immunity. J Immunol 190(6):2712–2719. https://doi.org/10.4049/jimmunol.1203082
Nemerow GR, Mold C, Schwend VK, Tollefson V, Cooper NR (1987) Identification of gp350 as the viral glycoprotein mediating attachment of Epstein-Barr virus (EBV) to the EBV/C3d receptor of B cells: sequence homology of gp350 and C3 complement fragment C3d. J Virol 61(5):1416–1420. https://doi.org/10.1128/jvi.61.5.1416-1420.1987
Nichols JW, Hill MW, Bangham AD, Deamer DW (1980) Measurement of net proton-hydroxyl permeability of large unilamellar liposomes with the fluorescent pH probe, 9-aminoacridine. Biochim Biophys Acta BBA Biomembr 596(3):393–403. https://doi.org/10.1016/0005-2736(80)90126-1
Nicolson GL (2013) Update of the 1972 Singer-Nicolson fluid-mosaic model of membrane structure. Discoveries (Craiova, Romania) 1(1):e3. https://doi.org/10.15190/d.2013.3
Nicolson GL, Ferreira de Mattos G (2021) A brief introduction to some aspects of the fluid-mosaic model of cell membrane structure and its importance in membrane lipid replacement. Membranes 11(12):947. https://doi.org/10.3390/membranes11120947
Niles WD, Cohen FS (1987) Video fluorescence microscopy studies of phospholipid vesicle fusion with a planar phospholipid membrane. Nature of membrane-membrane interactions and detection of release of contents. J Gen Physiol 90:703–735. https://doi.org/10.1085/jgp.90.5.703
Nir S, Wilschut J, Bentz J (1982) The rate of fusion of phospholipid vesicles and the role of bilayer curvature. Biochim Biophys Acta 688(1):275–278. https://doi.org/10.1016/0005-2736(82)90604-6
Orci L, Glick BS, Rothman JE (1986) A new type of coated vesicular carrier that appears not to contain clathrin: its possible role in protein transport within the Golgi stack. Cell 46:171–184. https://doi.org/10.1016/0092-8674(86)90734-8
Oswald AT, MacLeod CM, McCarty M (1944) Studies on the chemical nature of the substance inducing transformation of Pneumococcal types: induction of transformation by a deoxyribonucleic acid fraction isolated from Pneumococcus Type III. J Exp Med 79(2):137–158. https://doi.org/10.1084/jem.79.2.137
Panáková D, Sprong H, Marois E, Thiele C, Eaton S (2005) Lipoprotein particles are required for Hedgehog and Wingless signalling. Nature 435(7038):58–65. https://doi.org/10.1038/nature03504
Pap E, Pallinger E, Pásztói M, Falus AJ (2009) Highlights of a new type of intercellular communication: microvesicle-based information transfer. Inflamm Res 58:1–8. https://doi.org/10.1007/s00011-008-8210-7
Pasek M, Goto T, Gilbert W, Zink B, Schaller H, MacKay P, Leadbetter G, Murray K (1979) Hepatitis B virus genes and their expression in E. coli. Nature 282(5739):575–9. https://doi.org/10.1038/282575a0
Pathan M, Fonseka P, Chitti SV, Kang T, Sanwlani R, Van Deun J, Hendrix A, Mathivanan S (2019) Vesiclepedia 2019: a compendium of RNA, proteins, lipids and metabolites in extracellular vesicles. Nucleic Acids Res. https://doi.org/10.1093/nar/gky1029
Pearse BM (1976) Clathrin: a unique protein associated with intracellular transfer of membrane by coated vesicles. PNAS 73(4):1255–1259. https://doi.org/10.1073/pnas.73.4.1255
Pegtel DM, Gould SJ (2019) Exosomes. Annu Rev Biochem 88:487–514. https://doi.org/10.1146/ANNUREV-BIOCHEM-013118-111902
Peng L, Ryazantsev S, Sun R, Zhou ZH (2010) Three-dimensional visualization of gammaherpesvirus life cycle in host cells by electron tomography. Structure 18(1):47–58. https://doi.org/10.1016/j.str.2009.10.017
Perin MS, Fried VA, Mignery GA, Jahn R, Sudhof TC (1990) Phospholipid binding by a synaptic vesicle protein homologous to the regulatory region of protein kinase C. Nature 345:260–263. https://doi.org/10.1038/345260a0
Persoon CM, Moro A, Nassal JP, Farina M, Broeke JH, Arora S, Dominguez N, van Weering JR, Toonen RF, Verhage M (2018) Pool size estimations for dense-core vesicles in mammalian CNS neurons. EMBO J 37:e99672. https://doi.org/10.15252/embj.201899672
Peters KK, Carley WW, Palade GE (1985) Endothelial plasmalemmal vesicles have a characteristic striped bipolar surface structure. J Cell Biol 101(6):2233–2238. https://doi.org/10.1083/jcb.101.6.2233
Raposo G, Stoorvogel W (2013) Extracellular vesicles: exosomes, microvesicles, and friends. J Cell Biol 200(4):373–383. https://doi.org/10.1083/jcb.201211138
Rasmussen S, Bedau MA, Chen L, Deamer D, Krakauer DC, Packard NH, Stadler PF (eds) (2004) Protocells: bridging nonliving and living matter. MIT Press, Cambridge. https://doi.org/10.7551/mitpress/9780262182683.001.0001
Rice NR, Henderson LE, Sowder RC, Copeland TD, Oroszlan S, Edwards JF (1990) Synthesis and processing of the transmembrane envelope protein of equine infectious anemia virus. J Virol 64(8):3770–3778. https://doi.org/10.1128/jvi.64.8.3770-3778.1990
Roth TF, Keith R, Porter KR (1964) Yolk protein uptake in the oocyte of the mosquito Aedes aegytpi L. J Cell Biol 20(2):313–332. https://doi.org/10.1083/jcb.20.2.313
Rothberg KG, Heuser JE, Donzell WC, Ying Y-S, Glenney JR, Anderson RGW (1992) Caveolin, a protein component of caveolae membrane coats. Cell 68(4):673–682. https://doi.org/10.1016/0092-8674(92)90143-Z
Rowe T, Aridor M, McCaffery JM, Plutner H, Nuoffer C, Balch WE (1996) COPII vesicles derived from mammalian endoplasmic reticulum microsomes recruit COPI. J Cell Biol 135(4):895–911. https://doi.org/10.1083/jcb.135.4.895
Salmons T, Kuhn A, Wylie F, Schleich S, Rodriguez JR, Rodriguez D, Esteban M, Griffiths G, Locker JK (1997) Vaccinia virus membrane proteins p8 and p16 are cotranslationally inserted into the rough endoplasmic reticulum and retained in the intermediate compartment. J Virol 71(10):7404–7420. https://doi.org/10.1128/jvi.71.10.7404-7420.1997
Sambrook J, Rodgers L, White J, Gething MJ (1985) Lines of BPV-transformed murine cells that constitutively express influenza virus hemagglutinin. EMBO J 4(1):91–103
Sarkar DP, Morris SJ, Eidelman O, Zimmerberg J, Blumenthal R (1989) Initial stages of influenza hemagglutinin-induced cell fusion monitored simultaneously by two fluorescent events: cytoplasmic continuity and lipid mixing. J Cell Biol 109:113–122. https://doi.org/10.1083/Jcb.109.1.113
Sarmiento MA, Haffey M, Spear PG (1979) Membrane proteins specified by herpes simplex viruses. III. Role of glycoprotein VP7 (B2) in virion infectivity. J Virol 29(3):1149–1158. https://doi.org/10.1128/jvi.29.3.1149-1158.1979
Schauflinger M, Villinger C, Mertens T, Walther P, von Einem J (2013) Analysis of human cytomegalovirus secondary envelopment by advanced electron microscopy. Cell Microbiol 15(2):305–314. https://doi.org/10.1111/cmi.12077
Scheid A, Choppin PW (1974) Identification of biological activities of paramyxovirus glycoproteins. Activation of cell fusion, hemolysis, and infectivity by proteolytic cleavage of an inactive precursor protein of Sendai virus. Virology 57(2):475–490. https://doi.org/10.1016/0042-6822(74)90187-1
Scheu M, Komorowski K, Shen C, Salditt T (2021) A stalk fluid forming above the transition from the lamellar to the rhombohedral phase of lipid membranes. Eur Biophys J 50:265–278. https://doi.org/10.1007/s00249-020-01493-2
Schröder BA, Wrocklage C, Hasilik A, Saftig P (2010) The proteome of lysosomes. Proteomics 10:4053–4076. https://doi.org/10.1002/PMIC.201000196
Serrano-Heras G, Díaz-Maroto I, Castro-Robles B, Carrión B, Perona-Moratalla AB, Gracia J, Arteaga S, Hernández-Fernández F, García-García J, Ayo-Martín O, Segura T (2020) Isolation and quantification of blood apoptotic bodies, a non-invasive tool to evaluate apoptosis in patients with ischemic stroke and neurodegenerative diseases. Biol Proced Online 22(1):1–7. https://doi.org/10.1186/s12575-020-00130-8
Shangguan T, Alford D, Bentz J (1996) Influenza virus-liposomes lipid mixing is leaky and largely insensitive to the material properties of the target membrane. Biochemistry 35:4956–4965. https://doi.org/10.1021/bi9526903
Shangguan T, Siegel D, Lear J, Axelsen P, Alford D, Bentz J (1998) Morphological changes and fusogenic activity of influenza virus hemagglutinin. Biophys J 74:54–62. https://doi.org/10.1016/S0006-3495(98)77766-5
Sharpe HJ, Stevens TJ, Munro S (2010) A comprehensive comparison of transmembrane domains reveals organelle-specific properties. Cell 142(1):158–169. https://doi.org/10.1016/j.cell.2010.05.037
Shimada T, Takagi J, Ichino T, Shirakawa M, Hara-Nishimura I (2018) Plant vacuoles. Annu Rev Plant Biol 69:123–145. https://doi.org/10.1146/ANNUREV-ARPLANT-042817-040508
Shimshick EJ, Mcconnell HM (1973) Lateral phase separation in phospholipid membranes. Biochemistry 12(12):2351–2360. https://doi.org/10.1021/bi00736a026
Siegel DP (2008) The Gaussian curvature elastic energy of intermediates in membrane fusion. Biophys J 95(11):5200–5215. https://doi.org/10.1529/biophysj.108.140152
Silverstein SC, Marcus PI (1964) Early stages of newcastle disease virus-hela cell interaction: an electron microscopic study. Virology 23(3):370–380. https://doi.org/10.1016/0042-6822(64)90259-4
Simpson RJ, Jensen SS, Lim JWE (2008) Proteomic profiling of exosomes: current perspectives. Proteomics 8:4083–4099. https://doi.org/10.1002/PMIC.200800109
Singer SJ, Nicolson GL (1972) The fluid mosaic model of the structure of cell membranes. Science 175(4023):720–731. https://doi.org/10.1126/science.175.4023.720
Singh S, Mittal A (2016) Transmembrane domain lengths serve as signatures of organismal complexity and viral transport mechanisms. Sci Rep 6:22352. https://doi.org/10.1038/srep22352
Singh S, Ponnappan N, Verma A, Mittal A (2019) Osmotic tolerance of avian erythrocytes to complete hemolysis in solute free water. Sci Rep 9:7976. https://doi.org/10.1038/s41598-019-44487-7
Sirohi D, Chen Z, Sun L, Klose T, Pierson TC, Rossmann MG, Kuhn RJ (2016) The 3.8 Å resolution cryo-EM structure of Zika virus. Science 352(6284):467–470. https://doi.org/10.1126/science.aaf5316
Smith JJ, Aitchison JD (2013) Peroxisomes take shape. Nat Rev Mol Cell Biol 14(12):803–817. https://doi.org/10.1038/nrm3700
Sollner T, Whiteheart SW, Brunner M, Erdjument-Bromage H, Geromanos S, Tempst P, Rothman JE (1993) SNAP receptors implicated in vesicle targeting and fusion. Nature 362:318–324. https://doi.org/10.1038/362318a0
Spruce AE, Iwata A, White JM, Almers W (1989) Patch clamp studies of single cell-fusion events mediated by a viral fusion protein. Nature 342(6249):555–558. https://doi.org/10.1038/342555a0
Spustova K, Köksal ES, Ainla A, Gözen I (2021) Subcompartmentalization and pseudo-division of model protocells. Small 17:2005320. https://doi.org/10.1002/smll.202005320
Stanley WM (1944) The size of influenza virus. J Exp Med 79(3):267–283. https://doi.org/10.1084/jem.79.3.267
Stegmann T, Schoen P, Bron R, Wey J, Bartoldus I, Ortiz A, Nieva JL, Wilschut J (1993) Evaluation of viral membrane fusion assays. Comparison of the octadecylrhodamine dequenching assay with the pyrene excimer assay. Biochemistry 32(42):11330–11337. https://doi.org/10.1021/bi00093a009
Stegmann T, Bartoldus I, Zumbrunn J (1995) Influenza hemagglutinin-mediated membrane fusion: influence of receptor binding on the lag phase preceding fusion. Biochemistry 34:1825–1832. https://doi.org/10.1021/bi00006a002
Steinkühler J, De Tillieux P, Knorr RL, Lipowsky R, Dimova R (2018) Charged giant unilamellar vesicles prepared by electroformation exhibit nanotubes and transbilayer lipid asymmetry. Sci Rep 8(1):1–9. https://doi.org/10.1038/s41598-018-30286-z
Steinkühler J, Knorr RL, Zhao Z, Bhatia T, Bartelt SM, Wegner S, Dimova R, Lipowsky R (2020) Controlled division of cell-sized vesicles by low densities of membrane-bound proteins. Nat Commun 11(1):905. https://doi.org/10.1038/s41467-020-14696-0
Steinkühler J, Fonda P, Bhatia T, Zhao Z, Leomil FSC, Lipowsky R, Dimova R (2021) Superelasticity of plasma- and synthetic membranes resulting from coupling of membrane asymmetry, curvature, and lipid sorting. Adv Sci (Weinh) 8(21):e2102109. https://doi.org/10.1002/advs.202102109
Struck DK, Hoekstra D, Pagano RE (1981) Use of resonance energy transfer to monitor membrane fusion. Biochemistry 20(14):4093–4099. https://doi.org/10.1021/bi00517a023
Sullivan R (2015) Epididymosomes: a heterogeneous population of microvesicles with multiple functions in sperm maturation and storage. Asian J Androl 17(5):726–729. https://doi.org/10.4103/1008-682X.155255
Surma MA, Klose C, Klemm RW, Ejsing CS, Simons K (2011) Generic sorting of raft lipids into secretory vesicles in yeast. Traffic 12(9):1139–1147. https://doi.org/10.1111/j.1600-0854.2011.01221.x
Swanson J (2008) Shaping cups into phagosomes and macropinosomes. Nat Rev Mol Cell Biol 9:639–649. https://doi.org/10.1038/nrm2447
Szoka FC, Papahadjopoulos D (1978) Procedure for preparation of liposomes with large internal aqueous space and high capture by reverse phase evaporation. Proc Natl Acad Sci USA 75:4194–4198. https://doi.org/10.1073/pnas.75.9.4194
Tajima M, Nakajima H, Ito Y (1969) Electron microscopy of equine infectious anemia virus. J Virol 4(4):521–527. https://doi.org/10.1128/jvi.4.4.521-527.1969
Tanford C (1973) The hydrophobic effect: formation of micelles and biological membranes. Wiley, New York
Tanford C (1978) The hydrophobic effect and the organization of living matter. Science 200:1012–1018. https://doi.org/10.1126/science.653353
Terlecky SR, Fransen M (2000) How peroxisomes arise. Traffic 1(6):465–473. https://doi.org/10.1034/j.1600-0854.2000.010604.x
Tieleman DP, Bentz J (2002) Molecular dynamics simulation of the evolution of hydrophobic defects in one monolayer of a phosphatidylcholine bilayer: relevance for membrane fusion mechanisms. Biophys J 83(3):1501–1510. https://doi.org/10.1016/S0006-3495(02)73920-9
Tiollais P, Charnay P, Vyas GN (1981) Biology of hepatitis B virus. Science 213(4506):406–411. https://doi.org/10.1126/science.6264599
Tordo N, Poch O (1988) Structure of rabies virus. In: Campbell JB, Charlton KM (eds) Rabies. Developments in veterinary virology, vol 7. Springer, Boston, pp 25–45. https://doi.org/10.1007/978-1-4613-1755-5_2
Tristram-Nagle S (2015) Use of X-ray and neutron scattering methods with volume measurements to determine lipid bilayer structure and number of water molecules/lipid. Subcell Biochem 71:17–43. https://doi.org/10.1007/978-3-319-19060-0_2
Tse FW, Iwata A, Almers W (1993) Membrane flux through the pore formed by a fusogenic viral envelope protein during cell fusion. J Cell Biol 121(3):543–552. https://doi.org/10.1083/jcb.121.3.543
Tuller G, Nemec T, Hrastnik C, Daum G (1999) Lipid composition of subcellular membranes of an FY1679-derived haploid yeast wild-type strain grown on different carbon sources. Yeast 15(14):1555–1564. https://doi.org/10.1002/(SICI)1097-0061(199910)15:14%3C1555::AID-YEA479%3E3.0.CO;2-Z
Ueno M, Tanford C, Reynolds JA (1984) Phospholipid vesicle formation using nonionic detergents with low monomer solubility. Kinetic factors determine vesicle size and permeability. Biochemistry 23:3070–3076. https://doi.org/10.1021/bi00308a03
Van der Pol E, Böing AN, Harrison P, Sturk A, Nieuwland R (2012) Classification, functions, and clinical relevance of extracellular vesicles. Pharmacol Rev 64(3):676–705. https://doi.org/10.1124/pr.112.005983
Veatch S, Keller SL (2005) Miscibility phase diagrams of giant vesicles containing sphingomyelin. Phys Rev Lett 94:48101. https://doi.org/10.1103/PhysRevLett.94.148101
Veatch SL, Polozov IV, Gawrisch K, Keller SL (2004) Liquid domains in vesicles investigated by NMR and fluorescence microscopy. Biophys J 86(5):2910–2922. https://doi.org/10.1016/S0006-3495(04)74342-8
Vijayakrishnan S, Loney C, Jackson D, Suphamungmee W, Rixon FJ, Bhella D (2013) Cryotomography of budding influenza A virus reveals filaments with diverse morphologies that mostly do not bear a genome at their distal end. PLoS Pathog 9(6):e1003413. https://doi.org/10.1371/journal.ppat.1003413
Vincent JP, Magee T (2002) Argosomes: membrane fragments on the run. Trends Cell Biol 12(2):57–60. https://doi.org/10.1016/S0962-8924(01)02227-9
Volchkov VE, Feldmann H, Volchkova VA, Klenk HD (1998) Processing of the Ebola virus glycoprotein by the proprotein convertase furin. Proc Nat Acad Sci 95(10):5762–5767. https://doi.org/10.1073/pnas.95.10.5762
Walch-Solimena C, Takei K, Marek KL, Midyett K, Sudhof TC, De Camilli P, Jahn R (1993) Synaptotagmin: a membrane constituent of neuropeptide-containing large dense-core vesicles. J Neurosci 13(9):3895–3903. https://doi.org/10.1523/JNEUROSCI.13-09-03895.1993
Wang Q, Luo Y, Xie J, Dong C, Weng S, Ai H, Lü L, Yang X, Yu X, He J (2008) Identification of two novel membrane proteins from the Tiger frog virus (TFV). Virus Res 136(1–2):35–42. https://doi.org/10.1016/j.virusres.2008.04.013
Wang N, Shi X, Jiang L, Zhang S, Wang D, Tong P, Guo D, Fu L, Cui Y, Liu X, Arledge KC (2013) Structure of MERS-CoV spike receptor-binding domain complexed with human receptor DPP4. Cell Res 23(8):986–993. https://doi.org/10.1038/cr.2013.92
Watson J, Crick F (1953) Molecular structure of nucleic acids: a structure for deoxyribose nucleic acid. Nature 171:737–738. https://doi.org/10.1038/171737a0
Weber T, Zemelman BV, McNew JA, Westermann B, Gmachl M, Parlati F, Sollner TH, Rothman JE (1998) SNAREpins: minimal machinery for membrane fusion. Cell 92:759–772. https://doi.org/10.1016/S0092-8674(00)81404-X
Weis WI, Bruenger AT, Skehel JJ, Wiley DC (1990) Refinement of the influenza virus hemagglutinin by simulated annealing. J Mol Biol 212:737–761. https://doi.org/10.1016/0022-2836(90)90234-D
White J, Helenius A (1980) pH-dependent fusion between the Semliki Forest virus membrane and liposomes. Proc Natl Acad Sci USA 77:3273–3277. https://doi.org/10.1073/pnas.77.6.3273
White J, Matlin K, Helenius A (1981) Cell fusion by Semliki Forest, influenza, and vesicular stomatitis viruses. J Cell Biol 89(3):674–9. https://doi.org/10.1083/jcb.89.3.674
White JM, Delos SE, Brecher M, Schornberg K (2008) Structures and mechanisms of viral membrane fusion proteins: multiple variations on a common theme. Crit Rev Biochem Mol Biol 43(3):189–219. https://doi.org/10.1080/10409230802058320
Wild TF, Malvoisin ET, Buckland R (1991) Measles virus: both the haemagglutinin and fusion glycoproteins are required for fusion. J Gen Virol 72(2):439–442. https://doi.org/10.1099/0022-1317-72-2-439
Wilschut J, Duzgunes N, Fraley R, Papahadjopoulos D (1980) Studies on the mechanism of membrane fusion: kinetics of calcium ion induced fusion of phosphatidylserine vesicles followed by a new assay for mixing of aqueous vesicle contents. Biochemistry 19:6011–6021. https://doi.org/10.1021/bi00567a011
Wilson I, Skehel J, Wiley D (1981) Structure of the haemagglutinin membrane glycoprotein of influenza virus at 3 Å resolution. Nature 289:366–373. https://doi.org/10.1038/289366a0
Witkowska A, Heinz LP, Grubmüller H, Jahn R (2021) Tight docking of membranes before fusion represents a metastable state with unique properties. Nat Commun 12(1):3606. https://doi.org/10.1038/s41467-021-23722-8
Woese CR, Kandler O, Wheelis ML (1990) Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya. Proc Nat Acad Sci USA 87(12):4576–4579. https://doi.org/10.1073/pnas.87.12.4576
Wong JY, Majewski J, Seitz M, Park CK, Israelachvili JN, Smith GS (1999) Polymer-cushioned bilayers. I. A structural study of various preparation methods using neutron reflectometry. Biophys J 77(3):1445–1457. https://doi.org/10.1016/S0006-3495(99)76992-4
Wong JY, Park CK, Seitz M, Israelachvili J (1999) Polymer-cushioned bilayers. II. An investigation of interaction forces and fusion using the surface forces apparatus. Biophys J 77(3):1458–1468. https://doi.org/10.1016/S0006-3495(99)76993-6
Xiao C, Chipman PR, Battisti AJ, Bowman VD, Renesto P, Raoult D, Rossmann MG (2005) Cryo-electron microscopy of the giant Mimivirus. J Mol Biol 353(3):493–496. https://doi.org/10.1016/j.jmb.2005.08.060
Xiao C, Kuznetsov YG, Sun S, Hafenstein SL, Kostyuchenko VA, Chipman PR, Suzan-Monti M, Raoult D, McPherson A, Rossmann MG (2009) Structural studies of the giant mimivirus. PLoS Biol 7(4):e1000092. https://doi.org/10.1371/journal.pbio.1000092
Xu H, Ren D (2015) Lysosomal physiology. Annu Rev Physiol 77:57–80. https://doi.org/10.1146/annurev-physiol-021014-071649
Xu F, Yang Z, Wang L, Lee YL, Yang CC, Xiao SY, Xiao H, Wen L (2007) Morphological characterization of hantavirus HV114 by electron microscopy. Intervirology 50(3):166–172. https://doi.org/10.1159/000098959
Yang L, Huang HW (2002) Observation of a membrane fusion intermediate structure. Science 297(5588):1877–1879. https://doi.org/10.1126/science.1074354
Yang L, Huang HW (2003) A rhombohedral phase of lipid containing a membrane fusion intermediate structure. Biophys J 84(3):1808–1817. https://doi.org/10.1016/S0006-3495(03)74988-1
Yang ST, Kiessling V, Simmons JA, White JM, Tamm LK (2015) HIV gp41-mediated membrane fusion occurs at edges of cholesterol-rich lipid domains. Nat Chem Biol 11(6):424–431. https://doi.org/10.1038/nchembio.1800
Yu X, Qiao M, Atanasov I, Hu Z, Kato T, Liang TJ, Zhou ZH (2007) Cryo-electron microscopy and three-dimensional reconstructions of hepatitis C virus particles. Virology 367(1):126–134. https://doi.org/10.1016/j.virol.2007.05.038
Yu IM, Zhang W, Holdaway HA, Li L, Kostyuchenko VA, Chipman PR, Kuhn RJ, Rossmann MG, Chen J (2008) Structure of the immature dengue virus at low pH primes proteolytic maturation. Science 319(5871):1834–1837. https://doi.org/10.1126/science.1153264
Zaitseva E, Mittal A, Griffin DE, Chernomordik LV (2005) Class II fusion protein of alphaviruses drives membrane fusion through the same pathway as class I proteins. J Cell Biol 169:167–177. https://doi.org/10.1083/jcb.200412059
Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, Zhao X, Huang B, Shi W, Lu R, Niu P (2020) A novel coronavirus from patients with pneumonia in China, 2019. New Eng J Med 382(8):727–33. https://doi.org/10.1056/NEJMoa2001017
Zimmerberg J, Blumenthal R, Sarkar DP, Curran M, Morris SJ (1994) Restricted movement of lipid and aqueous dyes through pores formed by influenza hemagglutinin during cell fusion. J Cell Biol 127:1885–1894. https://doi.org/10.1083/jcb.127.6.1885
Acknowledgements
AC is grateful to IIT Delhi for financial support. AM appreciates the importance of several interactions since the turn of the century that have contributed to the formulation of his current thoughts on role of biological membranes in origins of life and evolution of biological systems. In particular, he is thankful for the mentorships (transformed into prized friendships) of Joe Bentz and Leonid (Leonya) Chernomordik. He is particularly indebted to Eugenia (Zhenya) Leikina for being an exceptional friend and teacher of membrane fusion assays in cell cultures (her heroic experiments in the cold-room wearing eskimo-jackets in late 1990s remain unparalleled till date) and cherishes his discussions with Kamran (Kam) Melikov, Elena Zaitseva (EZ), Helene Delanoe-Ayari, Corrine Ramos (Co), Mukesh Kumar, Samuel (Sam) Hess, B. Jayaram (Jay), Prashant Mishra, and MNV Ravikumar. He also acknowledges interactions/discussions with Ken Dill, Harma Ellens, Dave Siegel, Robert Blumenthal, Barry Lentz, Benjamin Podbilewicz, Judith White, Manju Bansal, Antoine Danchin, Ken Miller, Pierre Pontarotti, Gustavo Caetano-Anollés, Jeremy Lee, Michael O'Connor, Ed Dzialowski, and Shivanthi Anandan - each one of the above have some contribution to his current thoughts. He appreciates the super long-term-continued positivity/encouragement from Josh Zimmerberg, Fred Cohen, Ramaswamy Sarma, and Jennifer Lippincott-Schwartz. He also wishes to acknowledge the role of Albany Conversations (2011, 2015, 2019) and Gordon conferences on Molecular Membrane Biology (2003, 2005) in contributing to the development of Fig. 4 of this work - GC MMBs also anecdotally for the pleasure of hiking with Randy Schekman and Tom Rappaport, along with fun interactions with Sandy Schmid (beyond getting a proper drubbing at the pool table). Finally, and importantly, AM is grateful for academic autonomy at IIT Delhi, and for funding to (i) Kusuma Trust (UK) towards establishment of the Kusuma School of Biological Sciences at IIT Delhi, (ii) Department of Biotechnology, Govt. of India towards establishment of SCFBio as a Center of Excellence at IIT Delhi, and (iii) CDAC Pune toward managing the grant to SCFBio under the aegis of the National Supercomputing Mission, Govt. of India.
Author information
Authors and Affiliations
Contributions
AM conceived and designed the study. AC collected most of the data for Tables 2, 3, and 4. The first draft of the manuscript was written by AM. The manuscript was finalized by AM with inputs from AC. Both authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Mittal, A., Chauhan, A. Aspects of Biological Replication and Evolution Independent of the Central Dogma: Insights from Protein-Free Vesicular Transformations and Protein-Mediated Membrane Remodeling. J Membrane Biol 255, 185–209 (2022). https://doi.org/10.1007/s00232-022-00230-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00232-022-00230-4