Abstract
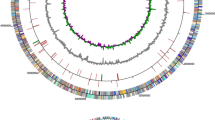
This study reports about the De novo whole genome sequencing and analysis of a bacterial isolate Streptomyces sp. Strain. RB7AG, isolated from the sediments of Chilika Lake, Odisha, India. The genome report in this paper highlights a size of 7,708,681 bp and a GC content of 72.3%. It also consists of 7274 coding sequence, 66 tRNA and 4 rRNA. Furthermore, carbohydrate active enzyme analysis revealed that the strain RB7AG has 127 glycoside hydrolase family genes, which is well known for hydrolysis of glycosidic bond in complex sugars. Thus, exploiting these microorganisms for the production of chitosan can be an appropriate waste disposal method of choice. Chitosan being an important biomolecule that has various industrial applications. Hence, the study also sought to improve the culture conditions for the Streptomyces sp. strain RB7AG for generation, recovery, and characterization of chitosan. Utilizing the isolate, various low-cost nitrogen sources, including peptone, yeast extract, ammonium chloride, urea along with pH, media, metal ions and surfactant were assessed for chitosan synthesis. In this context, traditional methods such as One Factor One Time are more time consuming and expensive too. The current work aims to establish a methodology to optimize the degradation of chitin by the chitinolytic Streptomyces sp. strain RB7AG, isolated from lake sediment for the production of chitosan. More than one factor was considered at the time of experiments, and the knowledge was integrated into Taguchi statistical design to determine the contribution of the most important factors required to achieve the desired end product i.e. chitosan. Highest chitosan production (2.188 µg/ml) was observed in MSM media, 1.0% NaCl (w/v), 0.5% Yeast extract, 1% Ca2+ and 0.1% Tween 80 at pH 9. The whole genome analysis of RB7AG would help in determining the mechanism involved in the breakdown activity.













Similar content being viewed by others
References
Adhikari HS, Yadav PN (2018) Anticancer activity of chitosan, chitosan derivatives, and their mechanism of action. Int J Biomater 2018:1–29
Aider M (2010) Chitosan application for active bio-based films production and potential in the food industry. LWT-Food Sci Technol 43:837–842
Ash K, Ramteke PW (2018) Optimization of extracellular alkaline protease production from Pseudomonas aeruginosa isolated from soil samples. Int J Agric Environ Biotechnol 11:187–194
Behera HT, Mojumdar A, Das SR, Jema S, Ray L (2020) Production of N-acetyl chitooligosaccharide by novel Streptomyces chilikensis strain RC1830 and its evaluation for anti-radical, anti-inflammatory, anti-proliferative and cell migration potential. Bioresour Technol Reports 11:100428
Biradar SP, Mehta MR, Mahajan HP, Bankhele RR, Hivrale AU (2022) Biological activities of chitosan-based nanomaterials. Role of chitosan and chitosan-based nanomaterials in plant sciences. Elsevier, Oxford
Chen J-K, Shen C-R, Liu C-L (2010) N-acetylglucosamine: production and applications. Mar Drugs 8:2493–2516
Chen H-J, Chang S-N, Tang C-W (2017) Application of the Taguchi method for optimizing the process parameters of producing lightweight aggregates by incorporating tile grinding sludge with reservoir sediments. Materials 10:1294
Collee JG, Miles RS, Watt B (1996) Tests for identification of bacteria. Mackie McCartney Prac Med Microbiol 14:131–149
Czitrom V (1999) One-factor-at-a-time versus designed experiments. Am Stat 53:126–131
Dasu VV, Panda T, Chidambaram M (2003) Determination of significant parameters for improved griseofulvin production in a batch bioreactor by Taguchi’s method. Process Biochem 38:877–880
Doroghazi JR, Albright JC, Goering AW, Kou-San Ju, Haines RR, Tchalukov KA, Labeda DP, Kelleher NL, Metcalf WW (2014) A roadmap for natural product discovery based on large-scale genomics and metabolomics. Nat Chem Biol 10:963–968
Felsenstein J (1985) Phylogenies and the comparative method. Am Nat 125:1–15
Ghaley BB, Hauggaard-Nielsen H, Høgh-Jensen H, Jensen ES (2005) Intercropping of wheat and pea as influenced by nitrogen fertilization. Nutr Cycl Agroecosyst 73:201–212
Gordon RE, Barnett DA, Handerhan JE, Pang CH-N (1974) Nocardia coeliaca, Nocardia autotrophica, and the nocardin strain. Int J Syst Evol Microbiol 24:54–63
Hsu SC, Lockwood JL (1975) Powdered chitin agar as a selective medium for enumeration of actinomycetes in water and soil. Appl Microbiol 29:422–426
Jahromi ST, Barzkar N (2018) Marine bacterial chitinase as sources of energy, eco-friendly agent, and industrial biocatalyst. Int J Biol Macromol 120:2147–2154
Jeon J-M, Thangamani R, Song E, Lee H-W, Lee H-W, Yang Y-H (2013) Media optimization of Corynebacterium glutamicum for succinate production under oxygen-deprived condition. J Microbiol Biotechnol 23:211–217
Kaur B, Kaur R (2013) Application of response surface methodology for optimizing arginine deiminase production medium for Enterococcus faecium sp. GR7. Sci World J. https://doi.org/10.1155/2013/892587
Khan R, Kaushik A, Solanki PR, Ansari AA, Pandey MK, Malhotra BD (2008) Zinc oxide nanoparticles-chitosan composite film for cholesterol biosensor. Anal Chim Acta 616:207–213
Kieser T, Bibb MJ, Buttner MJ, Chater KF, Hopwood DA (2000) Practical streptomyces genetics. John Innes Foundation, Norwich
Kuroiwa T, Izuta H, Nabetani H, Nakajima M, Sato S, Mukataka S, Ichikawa S, Kim S-K, Karadeniz F, Karadeniz F (2005) キチン・キトサン研究= Chitin and chitosan research 13 (2), 212, 2007–08-01. Carbohyd Polym 62:357–368
Lan W, Wang W, Zhimin Yu, Qin Y, Luan J, Li X (2016) Enhanced germination of barley (Hordeum vulgare L.) using chitooligosaccharide as an elicitor in seed priming to improve malt quality. Biotech Lett 38:1935–1940
Land M, Hauser L, Jun S-R, Nookaew I, Leuze MR, Ahn T-H, Karpinets T, Lund O, Kora G, Wassenaar T (2015) Insights from 20 years of bacterial genome sequencing. Funct Integr Genom 15:141–161
Lee BH (2008) Structure, function and applications of microbial β-galactosidase (lactase). Carbohydrate-active enzymes. Elsevier, New York
Linden JC, Stoner RJ, Knutson KW, Gardner-Hughes CA (2000) Organic disease control elicitors. Agro Food Ind Hi Tech 11:32–34
Lu H-D, Zhao H-Q, Wang K, Lv L-L (2011) Novel hyaluronic acid–chitosan nanoparticles as non-viral gene delivery vectors targeting osteoarthritis. Int J Pharm 420:358–365
Lv J, Lv X, Ma M, Oh D-H, Jiang Z, Fu X (2022) Chitin and chitin-based biomaterials: a review of advances in processing and food applications. Carbohyd Polym 22:120142
Mojumdar A, Upadhyay AK, Raina V, Ray L (2019) A simple and rapid colorimetric method for the estimation of chitosan produced by microbial degradation of chitin waste. J Microbiol Methods 158:66–70
Orzali L, Corsi B, Forni C, Riccioni L (2017) Chitosan in agriculture: a new challenge for managing plant disease. Biological activities and application of marine polysaccharides. Intech, London, pp 87–96
Othman SI, Alturki AM, Abu-Taweel GM, Altoom NG, Allam AA, Abdelmonem R (2021) Chitosan for biomedical applications, promising antidiabetic drug delivery system, and new diabetes mellitus treatment based on stem cell. Int J Biol Macromol 190:417–432
Panda BP, Ali M, Javed S (2007) Fermentation process optimization. Res J Microbiol 2:201–208
Pandey AR, Singh US, Momin M, Bhavsar C (2017) Chitosan: application in tissue engineering and skin grafting. J Polym Res 24:1–22
Passi A, Tibocha-Bonilla JD, Kumar M, Tec-Campos D, Zengler K, Zuniga C (2021) Genome-scale metabolic modeling enables in-depth understanding of big data. Metabolites 12:14
Peng W, Zhong J, Yang J, Ren Y, Tan Xu, Xiao S, Zhou J, Tan H (2014) The artificial neural network approach based on uniform design to optimize the fed-batch fermentation condition: application to the production of iturin A. Microb Cell Fact 13:1–10
Qin C, Yumin Du, Xiao L, Li Z, Gao X (2002) Enzymic preparation of water-soluble chitosan and their antitumor activity. Int J Biol Macromol 31:111–117
Rostami E (2020) Progresses in targeted drug delivery systems using chitosan nanoparticles in cancer therapy: a mini-review. J Drug Deliv Sci Technol 58:101813
Roy RK (2001) Design of experiments using the Taguchi approach: 16 steps to product and process improvement. Wiley, New York
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4(4):406–425
Samarasinghe RM, Kanwar RK, Kanwar JR (2014) The effect of oral administration of iron saturated-bovine lactoferrin encapsulated chitosan-nanocarriers on osteoarthritis. Biomaterials 35:7522–7534
Santos VP, Marques NSS, Maia PCSV, de Lima MAB, de OliveiraFranco L, de Campos-Takaki GM (2020) Seafood waste as attractive source of chitin and chitosan production and their applications. Int J Mol Sci 21:4290
Shirling EBT, Gottlieb D (1966) Methods for characterization of Streptomyces species. Int J Syst Bacteriol 16:313–340
Smibert RM (1994) Phenotypic characterization. Methods for general and molecular bacteriology.
Taguchi G (1986) Introduction to quality engineering: designing quality into products and processes
Tindall BJ, Sikorski J, Smibert RA, Krieg NR (2007) Phenotypic characterization and the principles of comparative systematics. Methods for general and molecular microbiology. ASM Press, Washington, DC, pp 330–393
Vickers NJ (2017) Animal communication: when i’m calling you, will you answer too? Curr Biol 27:R713–R715
Vijayaraghavan P, Arun A, Al-Dhabi NA, Vincent SGP, Arasu MV, Choi KC (2016) Novel Bacillus subtilis IND19 cell factory for the simultaneous production of carboxy methyl cellulase and protease using cow dung substrate in solid-substrate fermentation. Biotechnol Biofuels 9:1–13
Wang Y, Fang X, An F, Wang G, Zhang X (2011) Improvement of antibiotic activity of Xenorhabdus bovienii by medium optimization using response surface methodology. Microb Cell Fact 10:1–15
Zhang J, Gao N-f (2007) Application of response surface methodology in medium optimization for pyruvic acid production of Torulopsis glabrata TP19 in batch fermentation. J Zhejiang Univ Sci B 8:98–104
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
No conflict of interest exist between the authors.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Behera, S.S., Nivedita, S., Das, S. et al. Whole genome sequencing of a novel chitinolytic Streptomyces sp. RB7AG reveals it’s chitosan production potential: optimization of the process through Taguchi experimental design. 3 Biotech 13, 198 (2023). https://doi.org/10.1007/s13205-023-03613-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-023-03613-z