Abstract
Helicobacter pylori, one of the most common bacterial pathogens of humans, colonizes the gastric mucosa, where it appears to persist throughout the host's life unless the patient is treated. Colonization induces chronic gastric inflammation which can progress to a variety of diseases, ranging in severity from superficial gastritis and peptic ulcer to gastric cancer and mucosal-associated lymphoma1. Strain-specific genetic diversity has been proposed to be involved in the organism's ability to cause different diseases or even be beneficial to the infected host2,3 and to participate in the lifelong chronicity of infection4. Here we compare the complete genomic sequences of two unrelated H. pylori isolates. This is, to our knowledge, the first such genomic comparison. H. pylori was believed to exhibit a large degree of genomic and allelic diversity, but we find that the overall genomic organization, gene order and predicted proteomes (sets of proteins encoded by the genomes) of the two strains are quite similar. Between 6 to 7% of the genes are specific to each strain, with almost half of these genes being clustered in a single hypervariable region.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Cover, T. L. & Blaser, M. J. Helicobacter pylori and gastroduodenal disease. Annu. Rev. Med. 42, 135– 145 (1992).
Atherton, J. C. J, Peek, R. M. J, tham, K. T., Cover, T. L. & Blaser, M. J. Clinical and pathological importance of heterogeneity in vacA, the vacuolating cytotoxin gene of Helicobacter pylori. Gastroenterology 12, 92– 99 (1997).
Blaser, M. J. Not all Helicobacter pylori strains are created equal: should all be eliminated? Lancet 349, 1020– 1022 (1997).
Logan, R. P. H. & Berg, D. E. Genetic diversity of Helicobacter pylori. Lancet 348 1462–1463 (1996).
Tomb, J.-F. The complete genome sequence of the gastric pathogen Helicobacter pylori . Nature 388, 539–547 (1997).
Curnow, A. W. et al. Glu-tRNAGln amidotransferase: a novel heterotrimeric enzyme required for correct decoding of glutamine codons during translation. Proc. Natl Acad. Sci. USA 94, 11819– 11826 (1997).
Akopyants, N. S. et al. Analyses of the cag pathogenicity island of Helicobacter pylori. Mol. Microbiol. 28, 37– 53 (1998).
Censini, S. et al. cag, a pathogenecity island of Helicobacter pylori, encodes type I-specific and disease-associated virulence factors. Proc. Natl Acad. Sci. USA 93, 14648– 14653 (1996).
Akopyanz, N., Bukanov, N. O., Westblom, T. U. & Berg, D. E. PCR-based RFLP analysis of DNA sequence diversity in the gastric pathogen Helicobacter pylori. Nucleic Acids Res. 20, 6221–6225 (1992).
Han, J., Yu, E., Lee, I. & Lee, Y. Diversity among clinical isolates of Helicobacter pylori in Korea. Mol. Cells 7, 544–547 (1997).
Kansau, I. et al. Genotyping of Helicobacter pylori isolates by sequencing of PCR products and comparison with the RAPD technique. Res. Microbiol. 147, 661–669 ( 1996).
Ilver, D. et al. Helicobacter pylori adhesin binding fucosylated histo-blood group antigens revealed by retagging. Science 279, 373–377 (1998).
Lee, W. K. et al. Construction of a Helicobacter pylori–Escherichia coli shuttle vector for gene transfer in Helicobacter pylori. Appl. Environ. Microbiol. 63, 4866– 4871 (1997).
Jiang, Q., Hiratsuka, K. & Taylor, D. E. Variability of gene order in different Helicobacter pylori strains contributes to genome diversity. Mol. Microbiol. 20, 833–842 ( 1996).
Suerbaum, S., Brauer-Steppkes, T., Labigne, A., Cameron, B. & Drlica, K. Topoisomerase I of Helicobacter pylori: juxtaposition with a flagellin gene (flaB) and functional requriement of a fourth zinc finger motif. Gene 210 , 151–161 (1998).
Tatusov, R. L. et al. Metabolism and evolution of Haemophilus influenzae deduced from a whole-genome comparison with Escherichia coli. Curr. Biol. 6, 279–291 ( 1996).
Saunders, N. J., Peden, J. F., Hood, D. w. & Moxon, E. R. Simple sequence repeats in the Helicobacter pylori genome. Mol. Microbiol. 27, 1091–1098 (1998).
Sakagami, T. et al. Atrophic gastrities changes in both Helicobacter felis and Helicobacter pylori infected mice are host dependent and separate from antral gastritis. Gut 39, 639– 648 (1996).
Fock, K. M. et al. Seroprevalence of Helicobacter pylori infection. Gastroenterology 114, A596 (1998 ).
Smith, D. R. Complete genome sequence of Methanobacetrium thermoautotrophicum deltaH: functional analysis and comparative genomics. J. Bacteriol. 179, 7135–7155 (1997).
Acknowledgements
We thank T. Dorman, M. Rubenfield, C. Butler, B. Andrews and K. MacCormack for assistance in finalizing and editing sequence information. D.E.T. is a member of the Canadian Bacterial Diseases Network and a Scientist with the Alberta Heritage Foundation for Medical Research.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Alm, R., Ling, LS., Moir, D. et al. Genomic-sequence comparison of two unrelated isolates of the human gastric pathogen Helicobacter pylori. Nature 397, 176–180 (1999). https://doi.org/10.1038/16495
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/16495
This article is cited by
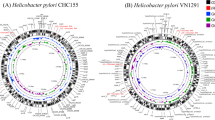
Comparative genomics of two Vietnamese Helicobacter pylori strains, CHC155 from a non-cardia gastric cancer patient and VN1291 from a duodenal ulcer patient
Scientific Reports (2023)
Helicobacter pylori infection
Nature Reviews Disease Primers (2023)
Trimer stability of Helicobacter pylori HtrA is regulated by a natural mutation in the protease domain
Medical Microbiology and Immunology (2023)
A graph-based approach for the visualisation and analysis of bacterial pangenomes
BMC Bioinformatics (2022)
Status of kinases in Epstein-Barr virus and Helicobacter pylori Coinfection in gastric Cancer cells
BMC Cancer (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.