Abstract
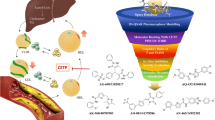
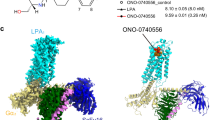
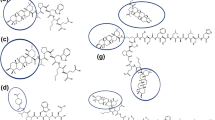
In this work, we designed three new ligands by conjugating cholesterol metabolites 3-hydroxy-5-cholestenoic acid (3-HC) and 3-oxo-4-cholestenoic acid (3-OC) and the natural tri-terpenoid betulinic acid with the tumor-targeting peptide YHWYGYTPQNVI. Molecular interactions with the unconjugated peptide and the conjugates were examined with three receptors that are commonly overexpressed in pancreatic adenocarcinoma cells using ligand docking and molecular dynamics. This study demonstrated the utility of the designed conjugates as a valuable scaffold for potentially targeting EGFR and LDLR receptors. Our results indicate that the conjugates showed strong binding affinities and formation of stable complexes with EGFR, while the unconjugated peptide, BT-peptide conjugate, an 3-HC-peptide conjugate showed the formation of fairly stable complexes with LDLR receptor. For EGFR, two receptor kinase domains were explored. Interactions with the N-terminal domain of CCKA-R were relatively weaker. For LDLR, binding occurred in the beta-propeller region. For the N-terminal fragment of CCKA-R, the conjugates induced significant conformational changes in the receptor. The molecular dynamic simulations for 100 ns demonstrate that BT-peptide conjugates and the unconjugated peptide had the highest binding and formed the most stable complexes with EGFR. RMSD and trajectory analyses indicate that these molecules transit to a dynamically stable configuration in most cases within 60 ns. NMA analysis indicated that amongst the conjugates that showed relatively higher interactions with the respective receptors, the highest potential for deformability was seen for the N-terminal-47 amino acid region of the CCKA-R receptor with and the lowest for the LDLR-receptor. Thus, the newly designed compounds may be evaluated in the future toward developing drug delivery materials for targeting tumor cells overexpressing LDLR or EGFR.
Graphical abstract













Similar content being viewed by others
Data availability
Associated data are included in the supplementary information. The manuscript will be shared on the faculty website and research square.
Code availability
None.
References
Siegel RL, Miller KD, Fuchs HE, Jemal A (2021) CA Cancer J Clin 71:7–33
Hellmann MD, Li BT, Chaft JE, Kris MG (2016) Ann Onc 27:1829–1835
Ventola CL (2017) Pharm Therapeutics 42:514–521
Mirjolet C, Papa AL, Créhange G, Raguin O, Seignez C, Paul C, Truc G, Maingon P, Millot N (2013) Radiother Oncol 108:136–142
Akhtar MJ, Ahamed M, Alhadlaq HA, Alrokayan SA, Kumar S (2014) Clin Chim Acta 436:78–92
Accardo A, Mansi R, Morisco A, Mangiapia G, Paduana L, Tesauro D, Radulescu A, Aurilio M, Aloj L, Arra C, Morelli G (2010) Mol Biosyst 6:878–887
Wang J, Liu W, Tu Q, Wang J, Song N, Zhang Y, Nie N, Wang J (2011) Biomacromol 12:228–234
Kang MH, Yoon HY, Choi YW (2017) Chapter 5- RIPL peptide as a novel cell penetrating and homing peptide: design, characterization and application to liposomal nanocarriers for hepsin-specific intracellular delivery, Nanostructures for Cancer Therapy, in Micro and Nanotechnologies 129–157
Choi YS, Lee JY, Suh JS, Lee SJ, Yang VC, Chung P, Park YJ (2011) Curr Pharm Biotechnol 12:1166–1182
Otte A, Jernmann E, Behe M, Goetze H, Bucher HC, Roser HW, Heppler A, Mueller-Brand J, Maecke HR (1997) Eur J Nucl Med 24:792–795
Romanov VI, Durand DB, Petrenko VA (2001) Prostate 47:239–251
Chau Y, Tan FE, Langer R (2004) Bioconjugate Chem 15:931–941
Feng S, Zou L, Ni Q, Zhang X, Li Q, Zheng L, Xie L, Li H, Huang D (2014) Cell Biochem Biophys 70:1913–1921
Acharya R, Chacko S, Bose P, Lapenna A, Pattanayak SP (2019) Sci Rep 9:15743
Oliveira-Cunha M, Newman WG, Siriwardena AK (2011) Cancers (Basel) 3:1513–1526
Schlessinger J (2000) Cell 103:211–225
Yarden Y (2001) Eur J Cancer 37:S3–S8
Henriksen L, Grandal MV, Knudsen SL, van Deurs B, Grøvdal LM (2013) PLoS One 8:e58148
Master AM, Sen Gupta A (2012) Nanomedicine (Lond) 7:1895–1906
Vasseur S, Guillaumond F (2015) Mol Cell Oncol 3: e1033586
Mayengbam SS, Singh A, Pillai AD, Bhat MK (2021) Transl Oncol 14:101043
Wank S (1998) Am J Physiol Gastrointest Liver Physiol 274:G607–G613
Weinberg DS, So C, Ruggeri B, Biswas S, Barber M, Waldman S (2000) Dig Dis Sci 45:538–543
Guillaumond F, Bidaut G, Ouaissi M, Servais S, Gouirand V, Olivares O, Lac S, Borge L, Roques J, Gayet O, Pinault M, Guimaraes C, Nigri J, Loncle C, Lavaut M, Garcia S, Tailleux A, Staels B, Calvo E, Tomasini R, Iovanna J, Vasseur S (2015) Proc Natl Acad Sci U S A 112:2473–2478
Sassi K, Nury T, Samadi M, Fennira FB, Vejux A, Lizard G (2021) Gliomas Chapter 6 not sure how to correctly cite when listed as chapter (97–120)
Radwan AA, Alanazi FK (2014) Saudi Pharm J 22:3–16
Ades A, Carvalho JP, Graziani SR, Amancio RF, Souen JS, Pinotti JA, Maranhão RC (2001) Gynecol Oncol 82:84–87
Ishimaru C, Yonezawa Y, Kuriyama I, Nishida M, Yoshida H, Mizushina Y (2008) Lipids 43:373–382
Sun L, Cao J, Chen K, Cheng L, Zhou C, Yan B, Qian W, Li J, Duan W, Ma J, Qi D, Wu E, Wang Z, Liu Q, Ma Q, Xu Q (2019) Int J Oncol 54:98–110
Fulda S (2008) Int J Mol Sci 9:1096–1107
Aiken C, Chen CH (2005) Trends Mol Med 11:31–36
Chowdhury AR, Mandal S, Mittra B, Sharma S, Mukhopadhyay S, Majumder HK (2002) Med Sci Monit 8:254–265
Schaap FG, Trauner M, Jansen PL (2014) Nat Rev Gastroenterol Hepatol 11:55–67
Hossein-Nejad-Ariani H, Althagafi E, Kaur K (2019) Sci Rep 9:2723
Master AM, Qi Y, Oleinick NL, Gupta AS (2012) Nanomedicine 8:655–664
Mickler FM, Möckl L, Ruthardt N, Ogris M, Wagner E, Bräuchle C (2012) Nano Lett 12:3417–3423
Abourbeh G, Shir A, Mishani E, Ogris M, Rödl W, Wagner E, Levitzki A (2012) IUBMB Life 64:324–330
Lin W, Chien W (2015) Journal Nanopart Res 17:349
The PyMOL Molecular Graphics System, Version 2.0 Schrodinger, LLC
Klamt A, Eckert F (2011) TmoleX3.1 COSMOlogic GmbH & amp; Co.: KG, Leverkusen, Germany
Eckert F, Klamt A (2002) AIChE J 48:369–385
Yu J, Zhou Y, Tanaka I, Yao M (2010) Bioinformatics 26:46–52
Yan C, Xu X, Zou X (2017) Modeling Peptide-Protein Interactions: Methods and Protocols. 3–9
Yan C, Zou X (2015) J Comput Chem 36:49–61
Trott O, Olson AJ (2010) J Comput Chem 31:455–461
Salentin S, Schreiber S, Haupt VJ, Adasme MF, Schroeder M (2015) Nucleic Acids Res 43:W443–W447
Bowers KJ, Chow E, Xu H, Dror RO, Eastwood MP, Gregersen BA, Klepeis JL, Kolossváry I, Moraes MA, Sacerdoti FD, Salmon JK, Shan Y, Shaw DE (2006) Proc. ACM/IEEE Conference on Supercomputing (SC06)
López-Blanco JR, Aliaga JI, Quintana-Ortí ES, Chacón P (2014) Nucleic Acids Res 42:W271–W276
Klamt A, Eckert F, Arlt W (2010) Annu Rev Chem Biomol Eng 1:101–122
Gonzalez-Miquel M, Massel M, DeSilva A, Palomar J, Rodriguez F, Brennecke JF (2014) J Phys Chem 118:11512–11522
Feng P, Che Y, Chen D (2020) Eur J Integr Med 33:101009
James K, Verkhivker GM (2014). PLoS ONE. https://doi.org/10.1371/journal.pone.0113488
Li Y, Li X, Ma W, Dong Z (2014) J Chem Theory Comput 10:3503–3511
Li Y, Li X, Dong Z (2015) J Comput Aided Mol Des 29:1045–1055
Park JH, Liu Y, Lemmon MA, Radhakrishnan R (2012) Biochem J 448:417–423
Xu G, Searle L, Hughes TV, Beck AK, Connolly PJ, Abad MC, Neeper MP, Struble GT, Springer BA, Emanuel SL, Gruniger RH, Pandey N, Adams M, Moreno-Mazza S, Fuentes-Pesquera AR (2008) Bioorg Med Chem Lett 18:3495–3499
Stamost J, Sliwkowski MX, Eigenbrot C (2002) J Biol Chem 277:46265–46272
Li Z, Zhao R, Wu X, Sun Y, Yao M, Li J, Xu Y, Gu J (2005) FASEB J 19:1978–1985
Ongarora B, Fontenot K, Hu X, Sehgal I, Satyanarayana-Jois S, Vicente M (2012) J Med Chem 55:3725–3738
Zhao Z, Xie L, Bourne PE (2019) J Chem Inf Model 59:453–462
Dong YW, Liao ML, Meng XL, Somero GN (2018) Proc Natl Acad Sci USA 115:1274–1279
Jin Z, Wang Y, Yu XF, Tan QQ, Liang SS, Li T, Zhang H, Shaw PC, Wang J, Hu C (2020) Comput Biol Chem 85:107241
Jiang X, Yu J, Zhou Z, Kongsted J, Song Y, Pannecouque C, Clercq ED, Kang D, Poongavanam V, Liu X, Zhang P (2019) Med Res Rev 39:1–45
Lobanov MY, Bogatyreva NS, Galzitskaya OV (2008) Mol Biol 42:623–628
Sammad FA, Suliman BA, Basha SH, Manivasagam T, Essa MM. (2016) PloS One 11: e0153999
Ali A, Banerjee S, Kamaal S, Usman M, Das N, Afzal M, Alarifi A, Sepay N, Roy P, Ahmad M (2021) RSC Adv 11:14362–14373
Beglova N, Blacklow SC (2005) Trends Biochem Sci 30:309–317
Wilson C, Wardell MR, Weisgraber KH, Mahley RW, Agard DA (1991) E Science 252:1817–1822
Andersen OM, Dagil R, Kragelund BB (2013) J Lipid Res 54:2763–2774
Pellegrini M, Mierke DF (1999) Biochemistry 38:14775–14783
Wank S (1995) Am J Physiol 269:G628–G646
Bahar I, Lezon TR, Bakan A, Shrivastava IH (2010) Chem Rev 110:1463–1497
Tirion MM, Ben-Avraham D (1993) J Mol Biol 230:186–195
Lopez-Blanco JR, Aliaga JI, Quintana-Orti ES, Chacon P (2014) Nucleic Acids Res 42:W271–W276
Author information
Authors and Affiliations
Contributions
Conception and design of study: I. A. Banerjee; Acquisition of data: M. M. Bashant, S. M. Mitchell, C. G. Lebedenko and L. R. Hart. Analysis and interpretation of data: I. A. Banerjee, M. M. Bashant and C. G. Lebedenko.
Corresponding author
Ethics declarations
Consent for publication
The authors give full consent for publication.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bashant, M.M., Mitchell, S.M., Hart, L.R. et al. In silico studies of interactions of peptide-conjugated cholesterol metabolites and betulinic acid with EGFR, LDR, and N-terminal fragment of CCKA receptors. J Mol Model 28, 16 (2022). https://doi.org/10.1007/s00894-021-05007-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-021-05007-5