Abstract
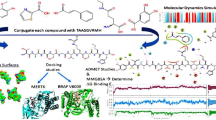
Peptide-based therapeutics have been gaining attention due to their ability to actively target tumor cells. Additionally, several varieties of nucleotide derivatives have been developed to reduce cell proliferation and induce apoptosis of tumor cells. In this work, we have developed novel peptide conjugates with newly designed purine analogs and pyrimidine derivatives and explored the binding interactions with the kinase domain of wild-type EGFR and its mutant EGFR [L858R/ T790M] which are known to be over-expressed in tumor cells. The peptides explored included WNWKV (derived from sea cucumber) and LARFFS, which in previous work was predicted to bind to Domain I of EGFR. Computational studies conducted to explore binding interactions include molecular docking studies, molecular dynamics simulations and MMGBSA to investigate the binding abilities and stability of the complexes. The results indicate that conjugation enhanced binding capabilities, particularly for the WNWKV conjugates. MMGBSA analysis revealed nearly twofold higher binding toward the T790M/L858R double mutant receptor. Several conjugates were shown to have strong and stable binding with both wild-type and mutant EGFR. As a proof of concept, we synthesized pyrimidine conjugates with both peptides and determined the KD values using SPR analysis. The results corroborated with the computational analyses. Additionally, cell viability and apoptosis studies with lung cancer cells expressing the wild-type and double mutant proteins revealed that the WNWKV conjugate showed greater potency than the LARFFS conjugate, while LARFFS peptide alone showed poor binding to the kinase domain. Thus, we have designed peptide conjugates that show potential for further laboratory studies for developing therapeutics for targeting the EGFR receptor and its mutant T790M/L858R.








Similar content being viewed by others
References
Vagace JM, Gervasini G (2011) Chemotherapy Toxicity in Patients with Acute Leukemia. In: Antica M (Eds), Acute Leukemia - The Scientist’s Perspective and Challenge. Intech Open. doi:https://doi.org/10.5772/841.
Sharma P, Hu-Lieskovan S, Wargo JA, Ribas A (2017) Primary, adaptive, and acquired resistance to cancer immunotherapy. Cell 168:707–723. https://doi.org/10.1016/j.cell.2017.01.017
Alfarouk KO, Stock C-M, Taylor S, Walsh M, Muddathir AK, Verduzco D, Bashir AH, Mohammed OY, Elhassan GO, Harguindey S, Reshkin SJ, Ibrahim ME, Rauch C (2015) Resistance to cancer chemotherapy: failure in drug response from ADME to P-GP. Cancer Cell Int. https://doi.org/10.1186/s12935-015-0221-1
Zhu X, Zhu H, Luo H, Zhang W, Shen Z, Hu X, Zhu X (2016) Molecular mechanisms of cisplatin resistance in cervical cancer. Drug Des Devel Ther 10:1885–1895. https://doi.org/10.2147/DDDT.S106412
Ma Y, Yu S, Ni S, Zhang B, Kung ACF, Gao J, Lu A, Zhang G (2021) Targeting strategies for enhancing paclitaxel specificity in chemotherapy. Front Cell Dev Biol. https://doi.org/10.3389/fcell.2021.626910
Xiao Y-F, Jie M-M, Li B-S, Li B-S, Hu C-J, Xie R, Tang B, Yang S-M (2015) Peptide-based treatment: a promising cancer therapy. J Immunol Res 2015:1–13. https://doi.org/10.1155/2015/761820
Feldmann G, Dhara S, Fendrich V, Bedja D, Beaty R, Mullendore M, Karikari C, Alvarez H, Lacobuzio-Donahue C, Jimeno A, Gabrielson KL, Matsui W, Maitra A (2007) Blockade of hedgehog signaling inhibits pancreatic cancer invasion and metastases: a new paradigm for combination therapy in solid cancers. Cancer Res 67:2187–2196. https://doi.org/10.1158/0008-5472.CAN-06-3281
Cheetham AG, Keith D, Zhang P, Lin R, Su H, Cui H (2016) Targeting tumors with small molecule peptides. Curr Drug Targets 16:489–508. https://doi.org/10.2174/1568009616666151130214646
Akhtar MJ, Ahamed M, Alhadlaq HA, Alrokayan SA, Kumar S (2014) Targeted anticancer therapy: overexpressed receptors and nanotechnology. Clin Chim Acta 436:78–92. https://doi.org/10.1016/j.cca.2014.05.004
Murayama O, Nishida H, Sekiguchi K (1996) Novel peptide ligands for Integrin α6β1 selected from a phage display library. J Biochem 120:445–451. https://doi.org/10.1093/oxfordjournals.jbchem.a021431
Chanda N, Kattumuri V, Shukla R, Zambre A, Katti K, Upendran A, Kulkarni RR, Kan P, Fent GM, Casteel SW, Smith CJ, Boote E, Robertson JD, Cutler C, Lever JR, Katti KV, Kannan R (2010) Bombesin functionalized gold nanoparticles show in vitro and in vivo cancer receptor specificity. Proc Natl Acad Sci USA 107:8760–8765. https://doi.org/10.1073/pnas.1002143107
He R, Finan B, Mayer JP, DiMarchi RD (2019) Peptide conjugates with small molecules designed to enhance efficacy and safety. Molecules 24:1855. https://doi.org/10.3390/molecules24101855
Li Y, Zheng X, Gong M, Zhang J (2016) Delivery of a peptide-drug conjugate targeting the blood brain barrier improved the efficacy of paclitaxel against glioma. Oncotarget 7:79401–79407. https://doi.org/10.18632/oncotarget.12708
Tesauro D, Accardo A, Aloj L, Aurilio M, Morelli G, Tesauro D (2014) Receptor binding peptides for target-selective delivery of nanoparticles encapsulated drugs. Int J Nanomed 9:1537–1557. https://doi.org/10.2147/IJN.S53593
Sigismund S, Avanzato D, Lanzetti L (2017) Emerging functions of the EGFR in cancer. Mol Oncol 12:3–20. https://doi.org/10.1002/1878-0261.12155
Kim JW, Kim YT, Kim DK, Song CH, Lee JW (1996) Expression of epidermal growth factor receptor in carcinoma of the cervix. Gynecol Oncol 60:283–287. https://doi.org/10.1006/gyno.1996.0039
Rimawi MF, Shetty PB, Weiss HL, Schiff R, Osborne CK, Channess GC, Elledge RM (2010) Epidermal growth factor receptor expression in breast cancer association with biologic phenotype and clinical outcomes. Cancer 116:1234–1242. https://doi.org/10.1002/cncr.24816
Sarabipour S (2017) Parallels and distinctions in FGFR, VEGFR, and EGFR mechanisms of transmembrane signaling. Biochemistry 56:3159–3173. https://doi.org/10.1021/acs.biochem.7b00399
Bethune G, Bethune D, Ridgway N, Xu Z (2010) Epidermal growth factor receptor (EGFR) in lung cancer: an overview and update. J Thorac Dis 2:48–51
Li AR, Chitale D, Riely GJ, Pao W, Miller VA, Zakowski MF, Rusch V, Kris MG, Ladanyi M (2008) EGFR mutations in lung adenocarcinomas. J Mol Diagn 10:242–248. https://doi.org/10.2353/jmoldx.2008.070178
Toyooka S, Kiura K, Mitsudomi T (2005) EGFR mutation and response of lung cancer to gefitinib. N Engl J Med 352:2136–2136. https://doi.org/10.1056/NEJM200505193522019
Grabe T, Lategahn J, Rauh D (2018) C797S resistance: The undruggable EGFR mutation in non-small cell lung cancer? ACS Med Chem Lett 9:779–782. https://doi.org/10.1021/acsmedchemlett.8b00314
Zhang T, Wan B, Zhao Y, Li C, Liu H, Lv T, Zhan P, Song Y (2019) Treatment of uncommon EGFR mutations in non-small cell lung cancer: new evidence and treatment. Transl Lung Cancer Res 8:302–316. https://doi.org/10.2137/tlcr.2019.04.12
Yan F, Liu X, Zhang S, Su J, Zhang Q, Chen J (2018) Effect of double mutations T790M/L858R on conformation and drug-resistant mechanism of epidermal growth factor receptor explored by molecular dynamics simulations. RSC Adv 8:39797–39810. https://doi.org/10.1039/C8RA06844E
Tamirat MZ, Koivu M, Elenius K, Johnson MS (2019) Structural characterization of EGFR exon 19 deletion mutation using molecular dynamics simulation. PLoS ONE. https://doi.org/10.1371/journal.pone.0222814
Suda K, Onozato R, Yatabe Y, Mitsudomi T (2009) EGFR T790M mutation: a double role in lung cancer cell survival? J Thorac Oncol 4:1–4. https://doi.org/10.1097/JTO.0b013e3181913c9f
Yun C-H, Boggon TJ, Li Y, Woo MS, Greulich H, Meyerson M, Eck MJ (2007) Structures of lung cancer-derived EGFR mutants and inhibitor complexes: mechanism of activation and insights into differential inhibitor sensitivity. Cancer Cell 11:217–227. https://doi.org/10.1016/j.ccr.2006.12.017
Thomson RJ, Moshirfar M, Ronquillo Y (2023) Tyrosine kinase inhibitors. StatPearls Publishing. https://www.ncbi.nlm.nih.gov/books/NBK563322/. Accessed 10 Jul 2023
Zhao Z, Xie L, Bourne PE (2018) Structural insights into characterizing binding sites in epidermal growth factor receptor kinase mutants. J Chem Inf Model 59:453–462. https://doi.org/10.1021/acs.jcim.8b00458
Jänne PA, Yang JC-H, Kim D-W, Planchard D, Ohe Y, Ramalingam SS, Ahn M-J, Kim S-W, Su W-C, Horn L, Haggstrom D, Felip E, Kim J-H, Frewer P, Cantarini M, Brown KH et al (2015) AZD9291 in EGFR inhibitor–resistant non–small-cell lung cancer. N Engl J Med 372:1689–1699. https://doi.org/10.1056/NEJMoa1411817
Singh PK, Silakari O (2018) Molecular dynamics guided development of indole based dual inhibitors of EGFR (T790M) and c-MET. Bioorg Chem 79:163–170. https://doi.org/10.1016/j.bioorg.2018.04.001
Ayati A, Moghimi S, Toolabi M, Foroumadi A (2021) Pyrimidine-based EGFR TK inhibitors in targeted cancer therapy. Eur J Med Chem 221:113523. https://doi.org/10.1016/j.ejmech.2021.113523
Romu AA, Lei Z, Zhou B, Chen Z-S, Korlipara V (2017) Design, synthesis and biological evaluation of WZ4002 analogues as EGFR inhibitors. Bioorg Med Chem 27:4832–4837. https://doi.org/10.1016/j.bmcl.2017.09.048
Sogabe S, Kawakita Y, Igaki S, Iwata H, Miki H, Cary DR, Takagi T, Takagi S, Ohta Y, Ishikawa T (2012) Structure-based approach for the discovery of pyrrolo[3,2-d] pyrimidine-based EGFR T790M/L858R mutant inhibitors. ACS Med Chem Lett 4:201–205. https://doi.org/10.1021/ml300327z
Tiefenbacher A, Pirker R (2017) EGFR tyrosine kinase inhibitors as first-line therapy in advanced EGFR mutation-positive non-small cell lung cancer: Strategies to improve clinical outcome. J Thorac Dis 9:4208–4211. https://doi.org/10.21037/jtd.2017.10.02
Wood ER, Truesdale AT, McDonald OB, Yuan D, Hassell A, Dickerson SH, Ellis B, Pennisi C, Horne E, Lackey K, Alligood KJ, Rusnak DW, Gilmer TM, Shewchuk L (2004) A unique structure for epidermal growth factor receptor bound to GW572016 (lapatinib). Cancer Res 64:6652–6659. https://doi.org/10.1158/0008-5472.CAN-04-1168
Sayed MT, Halim PA, El-Ansary AK, Hassan RA (2023) Design, synthesis, anticancer evaluation, and in silico studies of some thieno [2,3-d]pyrimidine derivatives as EGFR inhibitors. Drug Dev Res. https://doi.org/10.1002/ddr.22088
Furman O, Zaporozhets A, Tobi D, Bazylevich A, Firer MA, Patsenker L, Gellerman G, Lubin BCR (2022) Novel cyclic peptides for targeting EGFR and EGRVIII mutation for drug delivery. Pharmaceutics 14:1505. https://doi.org/10.3390/pharmaceutics14071505
Gavriil E-S, Doukatas A, Karampelas T, Myrianthopoulos V, Dimitrakis S, Mikros E, Marakos P, Tamvakopoulos C, Pouli N (2019) Design, synthesis and biological evaluation of novel substituted purine isosteres as EGFR kinase inhibitors, with promising pharmacokinetic profile and in vivo efficacy. Eur J Med Chem 176:393–409. https://doi.org/10.1016/j.ejmech.2019.05.029
Zhou H, Fu H, Liu H, Shao X, Cai W (2022) Uncovering the mechanism of drug resistance caused by the T790M mutation in EGFR kinase from absolute binding free energy calculations. Front Mol Biosci. https://doi.org/10.3389/fmolb.2022.922839
Zhang Y, He S, Bonneil É, Simpson BK (2020) Generation of antioxidative peptides from Atlantic sea cucumber using alcalase versus trypsin: in vitro activity, de novo sequencing, and in silico docking for in vivo function prediction. Food Chem 306:125581. https://doi.org/10.1016/j.foodchem.2019.125581
Yaghoubzadeh Z, Peyravii Ghadikolaii F, Kaboosi H, Safari R, Fattahi E (2019) Antioxidant activity and anticancer effect of bioactive peptides from rainbow trout (Oncorhynchus mykiss) skin hydrolysate. Int J Pept Res 26:625–632. https://doi.org/10.1007/s10989-019-09869-5
Song S, Liu D, Peng J, Deng H, Guo Y, Xu LX, Miller AD, Xu Y (2009) Novel peptide ligand directs liposomes toward EGF-R high-expressing cancer cells in vitro and in vivo. FASEB J 23:1396–1404. https://doi.org/10.1096/fj.08-117002
Tognolini M, Incerti M, Pala D, Russo S, Castelli R, Hassan-Mohamed I, Giorgio C, Lodola A (2013) Target hopping as a useful tool for the identification of novel EPHA2 protein-protein antagonists. Chem Med Chem 9:67–72. https://doi.org/10.1002/cmdc.201300305
Ghanem H, Kantarjian H, Ohanian M, Jabbour E (2012) The role of clofarabine in acute myeloid leukemia. Leuk Lymphoma 54:688–698. https://doi.org/10.3109/10428194.2012.726722
Daly MB, Roth ME, Bonnac L, Maldonado JO, Xie J, Clouser CL, Patterson SE, Kim B, Mansky LM (2016) Dual anti-HIV mechanism of clofarabine. Retrovirology 13:20. https://doi.org/10.1186/s12977-016-0254-0
Karmacharya U, Guragain D, Chaudhary P, Jee J-G, Kim J-A, Jeong B-S (2021) Novel pyridine Bioisostere of cabozantinib as a potent C-met kinase inhibitor: synthesis and anti-tumor activity against hepatocellular carcinoma. Int J Mol Sci 22:9685. https://doi.org/10.3390/ijms22189685
Xiao Q, Qu R, Gao D, Yan Q, Tong L, Zhang W, Ding J, Xie H, Li Y (2016) Discovery of 5-(methylthio) pyrimidine derivatives as L858R/T790M mutant selective epidermal growth factor receptor (EGFR) inhibitors. Bioorg Med Chem 24:2673–2680. https://doi.org/10.1016/j.bmc.2016.04.032
Tyagi A, Kapoor P, Kumar R, Chaudhary K, Gautam A, Raghava GPS (2013) In silico models for designing and discovering novel anticancer peptides. Sci Rep 3:2984. https://doi.org/10.1038/srep02984
Agarwal P, Bhagat D, Mahalwal M, Sharma N, Raghava GPS (2021) AntiCP 2.0: an updated model for predicting anticancer peptides. Brief Bioinform. https://doi.org/10.1093/bib/bbaa153
Schrödinger L, DeLano W (2020) PyMOL. Accessed http://www.pymol.org/pymol.
Balasubramani SG, Chen GP, Coriani S, Diedenhofen M, Frank MS, Franzke YJ, Furche F, Grotjahn R, Harding ME, Hattig C, Hellweg A, Helmich-Paris B, Holzer C, Hunair U, Kaupp M et al (2020) TURBOMOLE: modular program suite for ab initio quantum-chemical and condensed-matter simulations. J Chem Phys 152:184107. https://doi.org/10.1063/5.0004635
Mullins E, Oldland R, Liu YA, Wang S, Sandler SI, Chen C-C, Zwolak M, Seavey KC (2006) Sigma-profile database for using cosmo-based thermodynamic methods. Ind Eng Chem Res 45:4389–4415. https://doi.org/10.1021/ie060370h
Peng Y-H, Shiao H-Y, Tu C-H, Liu P-M, Hsu J, Amancha PK, Wu J-S, Coumar MS, Chen C-H, Wang S-Y, Lin W-H, Sun H-Y, Chao Y-S, Lyu P-C, Hsieh H-P, Wu S-Y (2013) Protein kinase inhibitor design by targeting the asp-phe-gly (DFG) motif: the role of the DFG motif in the design of epidermal growth factor receptor inhibitors. J Med Chem 56:3889–3903. https://doi.org/10.1021/jm400072p
Engelhardt H, Böse D, Petronczki M, Scharn D, Bader G, Baum A, Bergner A, Chong E, Döbel S, Egger G, Engelhardt C, Ettmayer P, Fuchs JE, Gerstberger T, Gonnella N, Grimm A, Grondal E, Haddad N, Hopfgartner B, Kousek R et al (2019) Start selective and rigidify: the discovery path toward a next generation of EGFR tyrosine kinase inhibitors. J Med Chem 62:10272–10293. https://doi.org/10.1021/acs.jmedchem.9b01169
Zardecki C, Dutta S, Goodsell DS, Lowe R, Voigt M, Burley SK (2021) PDB-101: educational resources supporting molecular explorations through biology and medicine. Protein Sci 31:129–140. https://doi.org/10.1002/pro.4200
Yu J, Zhou Y, Tanaka I, Yao M (2009) Roll: a new algorithm for the detection of protein pockets and cavities with a rolling probe sphere. Bioinformatics 26:46–52. https://doi.org/10.1093/bioinformatics/btp599
Halgren TA (2009) Identifying and characterizing binding sites and assessing druggability. J Chem Inf Model 49:377–389. https://doi.org/10.1021/ci800324m
Halgren TA (2007) New method for fast and accurate binding-site identification and analysis. Chem Biol Drug Des 69:146–148. https://doi.org/10.1111/j.1747-0285.2007.00483.x
Harris R, Olson AJ, Goodsell DS (2008) Automated prediction of ligand-binding sites in proteins. Proteins 70:1506–1517. https://doi.org/10.1002/prot.21645
Eberhardt J, Santos-Martins D, Tillack AF, Forli S (2021) AutoDock Vina 1.2.0: new docking methods, expanded force field, and python bindings. J Chem Inf Model 61:3891–3898. https://doi.org/10.1021/acs.jcim.1c00203
dos Santos KB, Guedes IA, Karl ALM, Dardenne L (2020) Highly Flexible Ligand Docking: benchmarking of the DockThor program on the LEADS-PEP protein-peptide dataset. J Chem Inf Model 60:667–683. https://doi.org/10.1021/acs.jcim.9b00905
Adasme MF, Linnemann KL, Bolz SN, Kaiser F, Salentin S, Haupt VJ, Schroeder M (2021) PLIP 2021: expanding the scope of the protein–ligand interaction profiler to DNA and RNA. Nucleic Acids Res 49:W530–W534. https://doi.org/10.1093/nar/gkab294
Bowers K, Chow E, Xu H, Dror RO, Eastwood MP, Gregersen BA, Klepeis JL, Kolossvary I, Moraes MA, Sacerdoti FD, Salmon JK, Shan Y, Shaw DE (2006) Scalable algorithms for molecular dynamics simulations on commodity clusters SC06: Proceedings of the 2006 ACM/IEEE conference on Supercomputing. https://doi.org/10.1145/1188455.1188544
Lu C, Wu C, Ghoreishi D, Chen W, Wang L, Damm W, Ross GA, Dahlgren MK, Russell E, Von Bargen CD, Abel R, Friesner RA, Harder ED (2021) OPLS4: Improving force field accuracy on challenging regimes of chemical space. J Chem Theory Comput 17:4291–4300. https://doi.org/10.1021/acs.jctc.1c00302
Jacobson MP, Pincus DL, Rapp CS, Day T, Honig B, Shaw DE (2004) A hierarchical approach to all-atom protein loop prediction. Proteins Struct Funct 55:351–367. https://doi.org/10.1002/prot.10613
Pattar SV, Adhoni SA, Kamanavalli CM, Kumbar SS (2020) In silico molecular docking studies and MM/GBSA analysis of coumarin-carbonodithioate hybrid derivatives divulge the anticancer potential against breast cancer. Beni-Suef Univ J Basic Appl Sci 9:36. https://doi.org/10.1186/s43088-020-00059-7
Li J, Abel R, Zhu K, Cao Y, Zhao S, Friesner RA (2011) The VSGB 2.0 model: a next generation energy model for high resolution protein structure modeling. Proteins 79:2794–2812. https://doi.org/10.1002/prot.23106
Xiong G, Wu Z, Yi J, Fu L, Yang Z, Hsieh C, Yin M, Zeng X, Wu C, Lu A, Chen X, Hou T, Cao D (2021) ADMETlab2.0: an integrated online platform for accurate and comprehensive predictions of ADMET properties. Nucleic Acids Res 49(W1):W5–W14. https://doi.org/10.1093/nar/gkab255
Nam K, Kimura T, Kishida A (2008) Controlling coupling reaction of EDC and NHS for preparation of collagen gels using ethanol/water co-solvents. Macromol Biosci 8:32–37. https://doi.org/10.1002/mabi.200700206
Yang HM, Teoh JY, Yim GK, Park Y, Kim YG, Kim J, Yoo D (2020) Label-free analysis of multivalent protein binding using bioresponsive nanogels and surface plasmon resonance (SPR). ACS Appl Mater Interfaces 12:5413–5419. https://doi.org/10.1021/acsami.9b17328
Harpaz D, Koh B, Marks RS, Seet RCS, Abdulalim I, Tok AIY (2019) Point-of-care surface plasmon resonance biosensor for stroke biomarkers NT-proBNP and S100β using a functionalized gold chip with specific antibody. Sensors 19:2533. https://doi.org/10.3390/s19112533
Minakata K, Takahashi F, Nara T, Hashimoto M, Tajima K, Murakami A, Nurwidya F, Yae S, Koizumi F, Moriyama H, Seyama K, Nishio K, Takahashi K (2012) Hypoxia induces gefitinib resistance in non-small-cell lung cancer with both mutant and wild-type epidermal growth factor receptors. Cancer Sci 103:1946–1954. https://doi.org/10.1111/j.1349-7006.2012.02408.x
Ngamwongsatit P, Banada PP, Panbangred W, Bhunia AK (2008) WST-1 based cell cytotoxicity assay as a substitute for MTT-based assay for rapid detection of toxigenic bacillus species using CHO cell line. J Microbiol Methods 73:211–215. https://doi.org/10.1016/j.mimet.2008.03.002
Chan A, Reiter R, Wiese S, Fertig G, Gold R (1998) Plasma membrane phospholipid asymmetry precedes DNA fragmentation in different apoptotic cell models. Histochem 110:553–558. https://doi.org/10.1007/s004180050317
Lakshmanan I, Batra S (2013) Protocol for apoptosis assay by flow cytometry using annexin v staining method. Bio protoc 3:374. https://doi.org/10.21769/bioprotoc.374
Rieger AM, Nelson KL, Konowalchuk JD, Barreda DR (2011) Modified annexin V/propidium iodide apoptosis assay for accurate assessment of cell death. J Vis Exp. https://doi.org/10.3791/2597
Catitti G, De Fabritiis S, Brocco D, Simeone P, De Bellis D, Vespa S, Veschi S, Lellis LD, Tinari N, Verginelli F, Marchisio M, Cama A, Patruno A, Lanuti P (2022) Flow cytometry detection of anthracycline-treated breast cancer cells: an optimized protocol. Curr Issues Mol Biol 45:164–174. https://doi.org/10.3390/cimb45010013
Klamt A (2011) The cosmo and cosmo-RS solvation models. Wiley Interdiscip Rev Comput Mol 1:699–709. https://doi.org/10.1002/wcms.56
Aissou M, Chemat-Djenni Z, Yara-Varón E, Fabiano-Tixier A-S, Chemat F (2017) Limonene as an agro-chemical building block for the synthesis and extraction of bioactive compounds. C R Chim 20:346–358. https://doi.org/10.1016/j.crci.2016.05.018
Modro AM, Modro TA (1999) The phosphoryl and the carbonyl group as hydrogen bond acceptors. Can J Chem 77:890–894
Osifová Z, Šála M, Dračínský M (2023) Hydrogen-bonding interactions of 8-substituted purine derivatives. ACS Omega. https://doi.org/10.1021/acsomega.3c03244
Milic M, Targos K, Tellez Chavez M, Thompson MAM, Jennings JJ, Franz AK (2021) NMR quantification of hydrogen-bond-accepting ability for organic molecules. J Org Chem 86:6031–6043. https://doi.org/10.1021/acs.joc.0c02876
Paul A, Thomas R (2022) Evidences for sulfur centered hydrogen bond with sulfur atoms as a donor in aromatic thiols and aliphatic thiols in aqueous solution. J Mol Liq 348:118078. https://doi.org/10.1016/j.molliq.2021.118078
Shan Y, Eastwood MP, Zhang X, Kim ET, Arkhipov A, Dror RO, Jumper J, Kuriyan J, Shaw DE (2012) Oncogenic mutations counteract intrinsic disorder in the EGFR kinase and promote receptor dimerization. Cell 149:860–870. https://doi.org/10.1016/j.cell.2012.02.063
Kaufman NE, Dhingra S, Jois SD, da Vicente M (2021) Molecular targeting of epidermal growth factor receptor (EGFR) and vascular endothelial growth factor receptor (VEGFR). Molecules 26:1076. https://doi.org/10.3390/molecules26041076
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31:455–461. https://doi.org/10.1002/jcc.21334
de Magalhães CS, Almeida DM, Barbosa HJC, Dardenne LE (2014) A dynamic niching genetic algorithm strategy for docking highly flexible ligands. Information Sci 289:206–224. https://doi.org/10.1016/j.ins.2014.08.002
de Magalhães CS, Barbosa HJC, Dardenne LE (2004) A Genetic algorithm for the ligand-protein docking problem. Genetics Mol Biol 27:605–610
Guedes IA, Barreto AMS, Marinho D, Kremser E, Kuenemann MA, Sperandio O, Dardenne LE, Miteva MA (2021) New machine learning and physics-based scoring functions for drug discovery. Sci Rep 11:3198
Treiber DK, Shah NP (2013) Ins and Outs of kinase DFG motifs. Chem Biol 20:745–746. https://doi.org/10.1016/j.chembiol.2013.06.001
Al-Warhi T, El Kerdawy AM, Said MA, Albohy A, Elsayed ZM, Aljaeed N, Elkaeed EB, Eldehna WM, Abdel-Aziz HA, Abdelmoaz MA (2022) Novel 2-(5-Aryl-4,5-dihydropyrazol-1-yl)thiazol-4-one as EGFR inhibitors: synthesis, biological assessment and molecular docking insights. Drug Des Devel Ther 16:1457–1471. https://doi.org/10.2147/DDDT.S356988
Stamos J, Sliwkowski MX, Eigenbrot C (2002) Structure of the epidermal growth factor receptor kinase domain alone and in complex with a 4-anilinoquinazoline inhibitor. J Biol Chem 277:46265–46272. https://doi.org/10.1074/jbc.M207135200
Kumar A, Petri ET, Halmos B, Boggon TJ (2008) Structure and clinical relevance of the epidermal growth factor receptor in human cancer. J Clin Oncol 26:1742–1751. https://doi.org/10.1200/JCO.2007.12.1178
Ezelarab H, Ali T, Abbas S, Sayed AM, Beshr EA, Hassan HA (2023) New antiproliferative 3-substituted oxindoles inhibiting EGFR/VEGFR-2 and tubulin polymerization. Mol Divers. https://doi.org/10.1007/s11030-023-10603-z
Nawaz F, Alam O, Perwez A, Rizvi MA, Naim MJ, Siddiqui N, Pottoo FH, Jha M (2020) 3′-(4-(Benzyloxy)phenyl)-1′-phenyl-5-(heteroaryl/aryl)-3,4-dihydro-1′H,2H-[3,4′-bipyrazole]-2-carboxamides as EGFR kinase inhibitors: synthesis, anticancer evaluation, and molecular docking studies. Arch Pharm 353:1900262. https://doi.org/10.1002/ardp.201900262
Ohkura K, Tabata A, Uto Y, Hori H (2020) Effect of isomerization of TX-2036 derivatives on the interaction with tyrosine kinase domain of EGF receptor. Anticancer Res 40:4675–4680. https://doi.org/10.21873/anticanres.14466
Jura N, Shan Y, Cao X, Shaw DE, Kuriyan J (2009) Structural analysis of the catalytically inactive kinase domain of the human EGF receptor 3. Proc Natl Acad Sci USA 106:21608–21613. https://doi.org/10.1073/pnas.0912101106
Mirza Z, Schulten H-J, Farsi HM, Al-Maghrabi JA, Gari MA, Chaudhary AG, Abuzenadah AM, Al-Qahtani MH, Karim S (2015) Molecular interaction of a kinase inhibitor midostaurin with anticancer drug targets, S100A8 and EGFR: transcriptional profiling and molecular docking study for kidney cancer therapeutics. PLoS ONE 10:1–17. https://doi.org/10.1371/journal.pone.0119765
Zare S, Emami L, Faghih Z, Zargari F, Zeinab F, Khabnadideh S (2023) Design, synthesis, computational study and cytotoxic evaluation of some new quinazoline derivatives containing pyrimidine moiety. Sci Rep 13:14461. https://doi.org/10.1038/s41598-023-41530-6
Chan C, Gill GN (1996) Mutational analysis of the nucleotide binding site of the epidermal growth factor receptor and V-src protein-tyrosine kinases. J Biol Chem 271:22619–22623. https://doi.org/10.1074/jbc.271.37.22619
Yaqeen Alhaqq FG, Mahdi M, Dawood A (2021) Theoretical drug design, molecular docking and ADME study of new 1,3,4-oxadiazole derivatives: promising anticancer agents against both breast and lung cancers. Egypt J Chem 64:6269–6283. https://doi.org/10.21608/ejchem.2021.75663.3735
Lemmon MA, Schlessinger J, Ferguson KM (2014) The EGFR family: not so prototypical receptor tyrosine kinases. Cold Spring Harb perspect biol 6:a020768. https://doi.org/10.1101/cshperspect.a020768
Mirtavoos-mahyari H, Rismani E, Lotfabadi AS, Dezfouli AA, Sheikhy K, Dezfuli MM, Hesmatania J (2021) Primary erlotinib resistance in a patient with non-small cell lung cancer carrying simultaneous compound EGFR L718A, Q849H, and L858R mutations. Biomol Concepts 12:164–174. https://doi.org/10.1515/bmc-2021-0018
Ercan D, Choi HG, Yun C-H, Capelletti M, Xie T, Eck MJ, Gray NS, Jänne PA (2015) EGFR mutations and resistance to irreversible pyrimidine-based EGFR inhibitors. Clin Cancer Res 21:3913–3923. https://doi.org/10.1158/1078-0432.CCR-14-2789
Pawara R, Ahmad I, Surana S, Patel H (2021) Computational identification of 2,4-disubstituted amino-pyrimidines as L858R/T790M-EGFR double mutant inhibitors using pharmacophore mapping, molecular docking, binding free energy calculation, DFT study and molecular dynamic simulation. In Silico Pharmacol 9:54. https://doi.org/10.1007/s40203-021-00113-x
Egieyeh S, Egieyeh E, Malan S, Christofells A, Fielding B (2021) Computational drug repurposing strategy predicted peptide-based drugs that can potentially inhibit the interaction of SARS-CO-2 spike protein with its target (human ACE2). PLoS ONE 16(1):e0245258. https://doi.org/10.1371/journal.pone.0245258
Benson NC, Faggett V (2008) Dynameomics: large-scale assessment of native protein flexibility. Protein Sci 17:2038–2050. https://doi.org/10.1110/ps.037473.108
Kalita MM, Fischer WB (2016) Asymmetric dynamics of ion channel forming proteins—hepatitis C virus (HCV) P7 bundles. Biochim Biophys Acta 1858:1462–1470. https://doi.org/10.1016/j.bbamem.2016.04.004
Robichaux JP, Le X, Vijayan RSK, Hicks JK, Heeke S, Elamin YY, Lin HY, Udagawa H, Skoulidis F, Tran H, Vargese S, He J, Zhang F, Nilsson MB, Hu L, Poteete A, Rinsurongkawong W, Zhang X, Ren C, Liu X, Hong L, Raymond V, Fang B, Wang J, Ha MJ, Cross JB, Gray JE, Heymach JV (2021) Structure-based classification predicts drug response in EGFR-mutant NSCLC. Nature 597:732–737. https://doi.org/10.1038/s41586-021-03898-1
Sharma J, Bhardwaj VK, Singh R, Rajendran V, Purohit R, Kumar S (2021) An in-silico evaluation of different bioactive molecules of tea for their inhibition potency against non structural protein-15 of SARS-COV-2. Food Chem 346:128933. https://doi.org/10.1016/j.foodchem.2020.128933
Moonrin N, Songtawee N, Rattanabunyong S, Chunsrivirot S, Mokmak W, Tonsima S, Choowongkomon K (2015) Understanding the molecular basis of EGFR kinase domain/ MIG-6 peptide recognition complex using computational analysis. BMC Bioinformatics 16:103. https://doi.org/10.1186/s12859-015-0528-x
Jura N, Shan Y, Cao X, Shaw D, Kuriyan J (2009) Structural analysis of the catalytically inactive kinase domain of the human EGF receptor 3. PNAS USA 106:21608–21613. https://doi.org/10.1073/pnas.0912101106
Kumar CK, Das S, Subramanian PT, Murali P, Isaac AE, Ramanathan K, Balamurli MM, Chanda K (2022) In-silico molecular modelling, MM/GBSA binding free energy and molecular dynamics simulation study of novel Pyrido fused imidazo [4,5-c]quinolines as potential anti-tumor agents. Front Chem. https://doi.org/10.3389/fchem.2022.991369
Onufriev AV, Case DA (2019) Generalized Born Implicit Models for Biomolecules. Annu Rev Biophys 48:275–296. https://doi.org/10.1146/annurev-biophys-052118-115325
Avdeef A (2001) Physicochemical profiling (solubility, permeability and charge state). Curr Med Chem 1:277–351. https://doi.org/10.2174/1568026013395100
Benet LZ, Hosey CM, Ursu O, Oprea TI (2016) BDDCS, the rule of 5 and drugability. Adv Drug Deliv Rev 101:89–98. https://doi.org/10.1016/j.addr.2016.05.007
Mitcheson JS (2008) HERG potassium channels and the structural basis of drug-induced arrhythmias. Chem Res Toxicol 21:1005–1010. https://doi.org/10.1021/tx800035b
Vijay U, Gupta S, Mathur P, Suravajhala P, Bhatnagar P (2018) Microbial mutagenicity assay: Ames test. Bio protoc 8:e2763. https://doi.org/10.21769/BioProtoc.2763
Irvine JD, Takahashi L, Lockhart K, Cheong J, Tolan JW, Selick HE (1999) MDCK (Madin-Darby canine kidney) cells: a tool for membrane permeability screening. J Pharm Sci 88:28–33. https://doi.org/10.1021/js9803205
Sharom FJ (2011) The P-glycoprotein multidrug transporter. Essays in Biochem 50:161–178. https://doi.org/10.1042/bse0500161
Montanari F, Ecker GF (2015) Prediction of drug–ABC-transporter interaction—recent advances and future challenges. Adv Drug Deliv Rev 86:17–26. https://doi.org/10.1016/j.addr.2015.03.001
McDonnell AM, Dang CH (2013) Basic review of the cytochrome P450 system. J Adv Pract Oncol 4:263–268. https://doi.org/10.6004/jadpro.2013.4.4.7
Guttman Y, Kerem Z (2022) Dietary inhibitors of CYP3A4 are revealed using virtual screening by using a new deep-learning classifier. J Agric Food Chem 70:2752–2761. https://doi.org/10.1021/acs.jafc.2c00237
Zhou Y, Nevosadová L, Eliasson E, Lauschke VM (2023) Global distribution of functionally important CYP2C9 alleles and their inferred metabolic consequences. Hum Genomics 17:15. https://doi.org/10.1186/s40246-023-00461-z
Sordella R, Bell DW, Haber DA, Settleman J (2004) Gefitinib-sensitizing EGFR mutations in lung cancer activate anti-apoptotic pathways. Science 305:1163–1167. https://doi.org/10.1158/1535-7163.MCT-05-0325
Zhang H, Shao H, Golubovskaya VM, Chen H, Cance W, Adjei AA, Dy GK (2016) Efficacy of focal adhesion kinase inhibition in non-small cell lung cancer with oncogenically activated MAPK PATHWAYS. Br J Cancer 115:203–211. https://doi.org/10.1038/bjc.2016.190
Ichim TE, O’Heeron P, Kesari S (2018) Fibroblasts as a practical alternative to mesenchymal stem cells. J Translational Med 16:212. https://doi.org/10.1186/s12967-018-1536-1
Virakul S, Dalm VAS, Paridaens D, van den Bosch W, Hirankarn N, van Hagen P, Dik W (2014) The tyrosine kinase inhibitor dasatinib effectively blocks PDGF-induced orbital fibroblast activation. Graefes Arch Clin Exp Ophthalmol 252:1101–1109. https://doi.org/10.1007/s00417-014-2674-7
Wang M, Chi CA (2018) Molecular mechanism of action and potential biomarkers of growth inhibition of synergistic combination of afatinib and dasatinib against gefitinib-resistant non-small cell lung cancer cells. Oncotarget 9:16533–16546. https://doi.org/10.18632/oncotarget.24814
Wang X, Xue Q, Wu L, Wang B, Liang H (2018) Dasatinib promotes TRAIL mediated apoptosis by upregulating CHOP-dependent death receptor 5 in gastric cancer. FEBS Open Bio 8:732–742. https://doi.org/10.1002/2211-5463.12404
Acknowledgements
HH, BG, MM, and MB thank the Henry Luce foundation for the Clare Boothe Luce Scholarship program for financial support of this work. The authors thank Ms. Chau Anh Phan for her help with running some of the surface plasmon resonance experiments. IB thanks the Fordham University Research Office for their support. The authors acknowledge the Advanced Research Computing, Education Technologies and Research Computing department at Fordham University, a division of the Office of Information Technology for providing their assistance and access to research computing resources that have contributed to the results reported here.
Funding
This study was funded by Fordham University Research office. IB thanks NSF-MRI grant # 2117625 for the acquisition of the Fluorescence-Activated Cell Sorter instrument.
Author information
Authors and Affiliations
Contributions
HLH: contributed toward in silico modeling, interpretation of data, data curation (both laboratory studies and in silico studies), and writing the first draft of the manuscript. BGG: contributed toward in vitro studies, data analysis, and writing parts of the manuscript; MAB: contributed toward data curation, in silico modeling, and in vitro studies; MIR: contributed toward data curation (cell viability studies); MEM: contributed toward data curation (SPRs); CGL: performed some of the molecular dynamics studies; IAB: contributed toward conceptualization, supervision, data analysis, editing, and writing the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hunt, H.L., Goncalves, B.G., Biggs, M.A. et al. Design and investigation of interactions of novel peptide conjugates of purine and pyrimidine derivatives with EGFR and its mutant T790M/L858R: an in silico and laboratory study. Mol Divers (2024). https://doi.org/10.1007/s11030-023-10772-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11030-023-10772-x