Abstract
The current need for forest conservation and management has driven a rapid expansion of landscape genetics approach. This discipline combines tools from molecular genetics, landscape ecology and spatial statistics and is decisive for improving not only ecological knowledge but also for properly managing population genetic resources. This approach could be appropriate to sweet chestnut (Castanea sativa Mill.), a multipurpose species of great economic importance in the Mediterranean basin and a species considered to be a good model of integration between natural and human-driven distribution of diversity. Sixteen chestnut populations, covering the distribution range of the species in Spain, were analysed using seven microsatellite markers. Results revealed a high level of genetic diversity in Spanish chestnut populations, which in part followed a geographical pattern, although distribution was not homogeneous. Likewise, areas particularly rich in diversity were detected, facilitating the development of a hypothesis about the history of chestnut in Spain. In conclusion, these results provide valuable baseline data for more in-depth studies on chestnut landscape genetics that can contribute to its conservation.




Similar content being viewed by others
References
Adua M (1999) The sweet chestnut throughout history from the Miocene to the third millennium. Acta Hort 494:29–36
Allendorf FW, Hohenlohe PA, Luikart G (2010) Genomics and the future of conservation genetics. Nat Rev Genet 11:697–709
Allué JL (1990) Atlas fitoclimático de España. Monografías INIA, no 69. MAPA-INIA, Madrid, p 221
Booy G, Hendricks RJJ, Smulders MJM, van Groenendael JM, Vosman B (2000) Genetic diversity and the survival of populations. Plant Biol 2:379–395
Buck EJ, Russell K, Hadonou M, James CJ, Blakesley D (2003) Isolation and characterization of polymorphic microsatellites in European chestnut (Castanea sativa Mill.). Mol Ecol Notes 3:239–241
Columela LJM (1979) De Res Rustica. Siglo I d.C. Los doce libros de Agricultura. Editorial Iberia, Barcelona
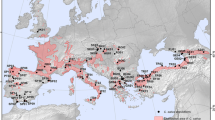
Conedera M, Krebs P, Tinner W, Pradella M, Torriani D (2004) The cultivation of Castanea sativa Miller in Europe, from its origin to its diffusion on a continental scale. Veg Hist Archaebot 13:161–179
Costa M, Morla C, Sainz H (1998) Los bosques ibéricos. Una interpretación paleobotánica. Editorial Planeta, Barcelona
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of cluster of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Fady B, Conord C (2010) Macroecological patterns of species and genetic diversity in vascular plants of the Mediterranean basin. Diversity Distrib 16:53–64
Fernandez-Lopez J, Monteagudo AB (2010) Genetic structure of wild Spanish populations of Castanea sativa as revealed by isozyme analysis. Forest Systems 19:156–169
Fineschi S, Taurchini D, Villani F, Vendarmin GG (2000) Chlororplast DNA polymorphism reveals little geograpgical structure in Castanea sativa Mill. (Fagaceae) throughout southern European countries. Mol Ecol 9:1495–1503
Frankman R, Ralls K (1998) Conservation biology: inbreeding leads to extinction. Nature 392:441–442
Garcia-Anton M, Morla C, Sainz H (1990) Consideraciones sobre la presencia de algunos vegetales relictos terciarios durante el cuaternario en la Península Ibérica. Bol R Soc Esp Hist Nat (Sec Biol) 86:95–105
Gomez A, Vendramin GG, Gonzalez-Martinez S, Alia R (2005) Genetic diversity and differentiation of two Mediterranean pines (Pinus halepensis Mill. and Pinus pinaster Ait.) along a latitudinal cline using chloroplast microsatellite markers. Drivers Distribution 11:257–263
Gomez-Sanz V, Blanco-Andray A, Sánchez-Palomares O, Rubio-Sánchez A, Elena-Roselló R, Graña-Domínguez D (2002) Autoecología de los castañares andaluces. Inv Agrar Sist Rec F 11:205–226
Goudet J (2002) FSTAT, a program to estimate and test gene diversities and fixation indices (version 2.9.3.2). Available from http://www.unil.ch/izea/softwares/fstat.html. Accessed 15 Sept 2010
Gupta PK, Balyan IS, Sharma PC, Ramesh B (1996) Microsatellite in plants—a new class of molecular markers. Curr Sci 70:45–54
Hedrick PW (2005) A standardised genetic differentiation measure. Evolution 59:1633–1638
Holderegger R, Wagner HH (2008) Landscape genetics. Bioscience 58:199–207
Huntley B, Kirks HJB (1983) An atlas of past and present pollen maps for Europe: 0–13,000 years ago. Cambridge University Press, Cambridge
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 14:1801–1806
Krebs P, Conedera M, Pradella M, Torriani D, Felber M, Tinner W (2004) Quaternary refugia of the sweet chestnut (Castanea sativa Miller): an extended palynological approach. Veg Hist Archaebot 13:145–160
Latta RG (2006) Integrating patterns across multiple genetic markers to infer spatial process. Landscape Ecol 21:809–820
Lauteri M, Monteverdi MC, Sansotta A, Cherubini M, Spaccino L, Villani F, Küçük M (1998) Adaptation to drought in European chestnut. Evidences from a hybrid zone and from controlled crosses between drought and wet adpated populations. Acta Hort 494:345–353
Manel S, Schwartz MK, Luikart G, Taberlet P (2003) Landscape genetics: combining landscape ecology and population genetics. Trends Ecol Evol 18:189–197
Manly BFJ (1985) The statistics of natural selection. Chapman and Hall, London
Marinoni D, Akkak A, Bounous G, Edwards KJ, Botta R (2003) Development and characterization of microsatellite markers in Castanea sativa (Mill.). Mol Breeding 11:127–136
Martín MA, Alvarez JB, Mattioni C, Cherubini M, Villani F, Martín LM (2009) Identification and characterisation of traditional chestnut varieties of southern Spain using morphological and simple sequence repeats SSR markers. Ann Appl Biol 154:389–398
Martín MA, Mattioni C, Cherubini M, Taurchini D, Villani F (2010) Neutral and adaptive genetic diversity in European chestnut populations by means of genomic and genic microsatellite markers. Tree Genet Genom 6:735–744
Mattioni C, Cherubini M, Micheli E, Villani F, Bucci G (2008) Role of domestication in shaping Castanea sativa genetic variation in Europe. Tree Genet Genomes 4:563–574
Miller MP (2005) Alleles in the Space (AIS): computer software for the joint analysis of interindividual spatial and genetic information. J Heredity 96:722–724
Monmonier M (1973) Maximun-differences barriers: an alternative numerical regionalization method. Geogr Anal 5:245–261
Namkoong G (2001) Forest genetics: pattern and complexity. Can J Forest Res 31:623–632
Nei M (1972) Genetic distance between populations. Amer Nat 106:283–392
Oddou-Muratorio S, Demuse-Munsh B, Pélissier R, Gouyon PH (2004) Impacts of gene flow and logging history on the local genetic structure of scattered tree species, Sorbus torminalis L. Crantz. Mol Ecol 13:3689–3702
Pautasso M (2009) Geographical genetics and the conservation of forest trees. Perspect Plant Ecol 11:157–189
Pereira-Lorenzo S, Costa R, Ramos-Cabrer A, Ribeiro C, da Silva M, Manzano G, Barreneche T (2010) Variation in grafted european chestnut and hybrids microsatellite reveals two main origins in the Iberian Peninsula. Tree Genet Genom 5:701–715
Petit RJ, Latouche-Hallé C, Pemonge MH, Kremer A (2002) Chloroplast DNA variation of oaks in France and the influence of forest fragmentation on genetic diversity. Forest Ecol Manag 156:115–129
Powell W, Machray GC, Provan J (1996) Polymorphism revealed by simple sequence repeats. Trends Plant Sci 1:215–222
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rivas-Martinez S (1987) Memoria del mapa de vegetación de España. 1:400.000. ICONA, MAPA, Madrid, p 110
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Saccheri I, Kuussaari M, Kankare M, Vikman P, Fortelius W, Hanski I (1998) Inbreeding and extintion in a butterfly metapopulation. Nature 392:491–494
Schneider S, Roessli D, Excoffier L (2000) Arlequin: a software for population genetics data analysis, version 3.1. Genetics and Biometry Laboratory, Department of Anthropology, University of Geneva
Shachak M, Boeken B, Groner E, Kadmon R, Lubin Y, Meron E, Neeman G, Perevolotsky A, Shkedy Y, Ungar ED (2008) Woody species as landscape modulators and their effect on biodiversity patterns. Bioscience 58:209–221
Sork VL, Smouse PE (2006) Genetic analysis of landscape connectivity in tree populations. Landscape Ecol 21:821–836
Storfer A, Murphy MA, Evans JS, Goldberg CS, Robinson S, Spear SF, Dezzani R, Delmelle E, Vierling L, Waits LP (2007) Putting the ‘landscape’ in landscape genetics. Heredity 98:128–142
Tautz D, Renz M (1984) Simple sequences are ubiquitous repetitive components of eukaryotic genomes. Nucleic Acids Res 12:4127–4138
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) Micro-Checker: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Vendramin GG, Scotti I, Ziegenhagen B (2004) Microsatellites in forest tree species: characteristics, identification and application. In: Kumar S, Fladung M (eds) Molecular genetics and breeding of forest trees. Haworth Press, New York, p 429
Villani F, Sansota A, Cherubini M, Cesaroni D, Sbordoni V (1999) Genetic structure of natural populations of Castanea sativa in Turkey: evidence of a hybrid zone. J Evol Biol 12:233–244
Watson DF (1992) Contouring: a guide to the analysis and display of spatial data. Pergamon Press, New York
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of populations structure. Evolution 38:1358–1370
Wright JP, Jones CG (2006) The concept of organisms as ecosystem engineers ten years on: progress, limitations, and challenges. Bioscience 56:203–209
Yeh FC, Yang RC, Boyle TBJ, Ye ZH, Mao JX (1997) Popgene ver 1.32. The user-friendly software for population genetic analysis. Molecular Biology and Biotechnology Center, University of Alberta, Canada
Acknowledgements
This research was supported by grants AGL2009-07931 and AGL2010-15147 from the Spanish Ministry of Science and Innovation and the European Regional Development Fund (FEDER) from the European Union. The first author is grateful to the Alfonso Martin Escudero Foundation for a postdoctoral fellowship and the hosting institute of the fellowship, Agroenvironmental and Forest Biology Institute from the Italian National Research Council.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A. Kremer
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table
Pairwise F ST estimates among all populations of chestnut included in this study. Significant pairwise comparisons (P < 0.05) are included in bold (DOC 56 kb)
Rights and permissions
About this article
Cite this article
Martín, M.A., Mattioni, C., Molina, J.R. et al. Landscape genetic structure of chestnut (Castanea sativa Mill.) in Spain. Tree Genetics & Genomes 8, 127–136 (2012). https://doi.org/10.1007/s11295-011-0427-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11295-011-0427-x