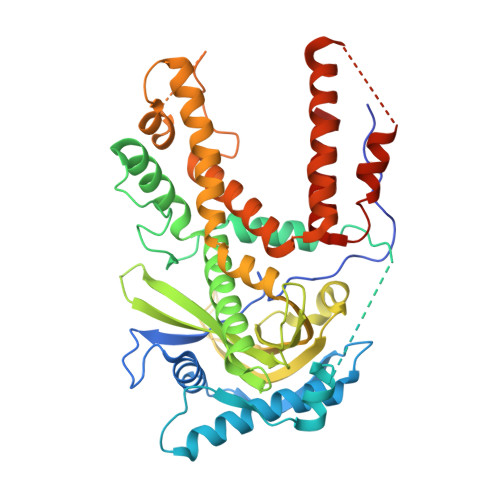
Fragment screening by STD NMR identifies novel site binders against influenza A virus polymerase PA
Pierce, P., Muruthi, M.M., Abendroth, J., Moen, S.O., Begley, D.W., Davies, D.R., Marathias, V.M., Staker, B.L., Myler, P.J., Lorimer, D.D., Edwards, T.E.To be published.