Abstract
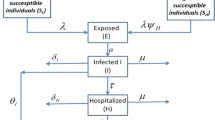
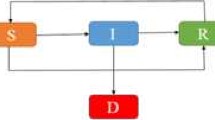
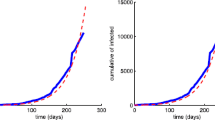
The deadly Ebola virus which spreads through human-to-human transmission via direct contact is a serious threat to the human population where the rate of death due to the Ebola virus infection is very high, with an average of approximately fifty percent, which has been seen to go as high as ninety percent in some outbreaks. In this paper, we attempt to understand the spreading behavior of Ebola virus disease (EVD) in the human population by developing a novel epidemic model with five classes susceptible-exposed-infectious-quarantine- infectious dead. We have used the very close real parametric values as per the data released by the World Health Organization (WHO). To minimize the spread of EVD, critical analysis is performed both in the presence and absence of control measures. The effect of quarantine on the infectious population is critically analyzed, and it is observed that quarantining the infectious population may play a vital role in controlling the spread of EVD. The reproduction number is obtained and the asymptotic stability is established both analytically and numerically. Extensive numerical simulations with examples and time series analysis are carried out under different control measures using MATLAB. Numerical simulations of our model and time series analysis reveal that the dead bodies are a major source of the spread of infection and thus our assumption that the deaths occurring among patients in the infectious or quarantined classes, lead to a transition into the dead (D) class, which contributes to the process of infection of susceptible members of the population is established. Once the dead bodies are buried, only they are assumed to be finally removed from the population. Introducing infectious dead class in our model is a novel idea that helped us to understand how we can minimize the transmission rate of EVD in the population.






Similar content being viewed by others
Data Availability
Enquiries about data availability should be directed to the authors.
Abbreviations
- S:
-
Susceptible population
- E:
-
Exposed population
- I:
-
Infectious population
- Q:
-
Quarantine population
- D:
-
Death population due to the Ebola virus before dead bodies are buried
- \(\mu\) :
-
Natural death rate i.e. death other than Ebola virus and the influx rate as well
- \(\delta_{1}\) :
-
Disease induced death rate for non-quarantined infectious individuals
- \(\delta_{2}\) :
-
Disease induced death rate for quarantined individuals
- \(\delta\) :
-
The rate at which dead bodies are buried
- \(\alpha\) :
-
The rate at which suspected cases (E) are quarantined
- \(\xi\) :
-
The rate at which confirmed cases (I) are quarantined
- \(\gamma\) :
-
The rate at which suspected cases (E) are confirmed
- \(\eta p\) :
-
The probability and rate of quarantine population to recovered and again joined the susceptible population
- \(\beta\) :
-
Per infectivity contact rate (from infectious patients to susceptible population)
- \(\beta_{D}\) :
-
Per infectivity contact rate (from dead bodies to susceptible population)
- \(\beta_{Q}\) :
-
Per infectivity contact rate (from quarantined patients to susceptible population mainly health workers)
References
https://www.who.int/news-room/fact-sheets/detail/ebola-virus-disease
https://www.who.int/emergencies/disease-outbreak-news/item/2021-DON312.
Meyers, L.A., Pourbohloul, B., Newman, M.E., Skowronski, D.M., Brunham, R.C.: Network theory and SARS: predicting outbreak diversity. J. Theor. Biol. 232(1), 71–81 (2005)
Kim, S., Lee, J., Jung, E.: Mathematical model of transmission dynamics and optimal control strategies for 2009 A/H1N1 influenza in the Republic of Korea. J. Theor. Biol. 412, 74–85 (2017)
Rizzo, A., Pedalino, B., Porfiri, M.: A network model for Ebola spreading. J. Theor. Biol. 394, 212–222 (2016)
Feng, Z., Zheng, Y., Hernandez-Ceron, N., Zhao, H., Glasser, J.W., Hill, A.N.: Mathematical models of Ebola—consequences of underlying assumptions. Math. Biosci. 277, 89–107 (2016)
Omame, A., Abbas, M., Onyenegecha, C.P.: Backward bifurcation and optimal control in a co-infection model for SARS-CoV-2 and ZIKV. Results Phys. 37, 105481 (2022)
Omame, A., Okuonghae, D.: A co-infection model for oncogenic human papillomavirus and tuberculosis with optimal control and Cost-Effectiveness Analysis. Optim. Control Appl. Methods 42(4), 1081–1101 (2021)
Omame, A., Abbas, M.: Modeling SARS-CoV-2 and HBV co-dynamics with optimal control. Physica A 615(1), 128607 (2023)
Rwezaura, H., Diagne, M.L., Omame, A., de Espindola, A.L., Tchuenche, J.M.: Mathematical modeling and optimal control of SARS-CoV-2 and tuberculosis co-infection: a case study of Indonesia. Model. Earth Syst. Environ. 8, 5493–5520 (2022)
Soufiane, B., Touaoula, T.M.: Global analysis of an infection age model with a class of nonlinear incidence rates. J. Math. Anal. Appl. 434(2), 1211–1239 (2016)
Bentout, S., Tridane, A., Djilali, S., Touaoula, T.M.: Age-structured modeling of COVID-19 epidemic in the USA. UAE and Algeria. Alex. Eng. J. 60(1), 401–411 (2021)
Bentout, S., Chekroun, A., Kuniya, T.: Parameter estimation and prediction for coronavirus disease outbreak 2019 (COVID-19) in Algeria. AIMS Public Health 7(2), 306–318 (2020)
Djilali, S., Bentout, S., Kumar, S., Touaoula, T.M.: Approximating the asymptomatic infectious cases of the COVID-19 disease in Algeria and India using a mathematical model. Int. J. Model., Simul., Sci. Comput. 13(4), 2250028 (2022)
Milligan, G.N., Barrett, A.D.: Vaccinology: An Essential Guide. Wiley Blackwell, Chichester, p. 310 (2015). ISBN 978-1-118-63652-7. OCLC 881386962
Fraser, C., Donnelly, C.A., Cauchemez, S., Hanage, W.P., Van Kerkhove, M.D., Hollingsworth, T.D., Griffin, J., Baggaley, R.F., Jenkins, H.E., Lyons, E.J., Jombart, T.: Pandemic potential of a strain of influenza A (H1N1): early findings. Science 324(5934), 1557–1561 (2009). https://doi.org/10.1126/science.1176062. Bibcode:2009Sci...324.1557F. PMC 3735127. PMID 19433588
Unless noted R0 values are from History and Epidemiology of Global Smallpox Eradication (Archived 2016–05–10 at the Wayback Machine), a module of the training course "Smallpox: Disease, Prevention, and Intervention". The CDC and the World Health Organization, 2001. Slide 17. This gives sources as "Modified from Epid Rev 1993;15: 265–302, Am J Prev Med 2001; 20 (4S): 88–153, MMWR 2000; 49 (SS-9); 27–38"
Guerra, F.M., Bolotin, S., Lim, G., Heffernan, J., Deeks, S.L., Li, Y., Crowcroft, N.S.: The basic reproduction number (R0) of measles: a systematic review. Lancet Infect. Diseases 17(12), e420–e428 (2017). https://doi.org/10.1016/S1473-3099(17)30307-9. ISSN 1473-3099. Accessed 18 Mar 2020
Ireland's Health Services. Health Care Worker Information (PDF). Accessed 27 Mar 2020
Kretzschmar, M., Teunis, P.F., Pebody, R.G.: Incidence and reproduction numbers of pertussis: estimates from serological and social contact data in five European countries. PLos Med. 7(6), e1000291 (2010). https://doi.org/10.1371/journal.pmed.1000291.PMC2889930.PMID20585374
Gani, R., Leach, S.: Transmission potential of smallpox in contemporary populations. Nature 414(6865), 748–751 (2001). https://doi.org/10.1038/414748a. ISSN 1476-4687. Accessed 18 Mar 2020
Wallinga, J., Teunis, P.: Different epidemic curves for severe acute respiratory syndrome reveal similar impacts of control measures. Am. J. Epidemiol. 160(6), 509–516 (2004). https://doi.org/10.1093/aje/kwh255. PMID 15353409. Archived from the original on October 6, 2007
Truelove, S.A., Keegan, L.T., Moss, W.J., Chaisson, L.H., Macher, E., Azman, A.S., Lessler, J.: Clinical and epidemiological aspects of diphtheria: a systematic review and pooled analysis. Clin Infect Diseases. https://doi.org/10.1093/cid/ciz808. Accessed 18 Mar 2020
Li, Q., Guan, X., Wu, P., Wang, X., Zhou, L., Tong, Y., et al.: Early transmission dynamics in Wuhan, China, of novel coronavirus-infected pneumonia. N. Engl. J. Med. (2020). https://doi.org/10.1056/NEJMoa2001316.PMID31995857
Riou, J., Althaus, C.L.: Pattern of early human-to-human transmission of Wuhan 2019 novel coronavirus (2019-nCoV), December 2019 to January 2020. Eurosurveillance (2020). https://doi.org/10.2807/15607917.ES.2020.25.4.2000058. PMC 7001239. PMID 32019669
Liu, T., Hu, J., Kang, M., Lin, L.: Time-varying transmission dynamics of Novel Coronavirus Pneumonia in China. bioRxiv (2020). https://doi.org/10.1101/2020.01.25.919787
Read, J.M., Bridgen, J.R.E., Cummings, D.A.T., et al.: Novel coronavirus 2019-nCoV: early estimation of epidemiological parameters and epidemic predictions. MedRxiv (2020). https://doi.org/10.1101/2020.01.23.20018549
Wu, J.T., Leung, K., Bushman, M., Kishore, N., Niehus, R., de Salazar, P.M., Cowling, B.J., Lipsitch, M., Leung, G.M.: Estimating clinical severity of COVID-19 from the transmission dynamics in Wuhan, China. Nat Med (2020). https://doi.org/10.1038/s41591-020-0822-7. ISSN 1546-170X
Ferguson, N.M., Cummings, D.A., Fraser, C., Cajka, J.C., Cooley, P.C., Burke, D.S.: Strategies for mitigating an influenza pandemic. Nature 442(7101), 448–452 (2006). https://doi.org/10.1038/nature04795.PMID16642006
Althaus, C.L.: Estimating the reproduction number of Ebola virus (EBOV) during the 2014 outbreak in West Africa. PLos Curr. (2014). arXiv:1408.3505. Bibcode:2014. https://doi.org/10.1371/currents.outbreaks.91afb5e0f279e7f29e7056095255b288. PMC 4169395. PMID 25642364
Coburn, B.J., Wagner, B.G. and Blower, S.: Modeling influenza epidemics and pandemics: insights into the future of swine flu (H1N1). BMC Med. (2009). https://doi.org/10.1186/1741-7015-7-30. PMID 19545404
Kucharski, A., Althaus, C.L.: The role of superspreading in Middle East respiratory syndrome coronavirus (MERS-CoV) transmission. Eurosurveillance 20(26), 14–18 (2015). https://doi.org/10.2807/1560-7917.ES2015.20.25.21167. PMID 26132768
Berge, T., Lubuma, J.M.-S., Moremedi, G.M., Morris, N., Kondera-Shava, R.: A simple mathematical model for Ebola in Africa. J. Biol. Dyn. 11(1), 42–74 (2017). https://doi.org/10.1080/17513758.2016.1229817
Funding
The authors have not disclosed any funding.
Author information
Authors and Affiliations
Contributions
AM wrote the Introduction part, developed the Mathematical Model. BKM established the stability and performed the simulation. AKK provided the examples and performed the analysis.
Corresponding author
Ethics declarations
Conflict of Interest
The authors have not disclosed any competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mishra, A., Mishra, B.K. & Keshri, A.K. Quarantine Model on the Transmission of Ebola Virus Disease in the Human Population with Infectious Dead Class. Int. J. Appl. Comput. Math 9, 133 (2023). https://doi.org/10.1007/s40819-023-01608-1
Accepted:
Published:
DOI: https://doi.org/10.1007/s40819-023-01608-1