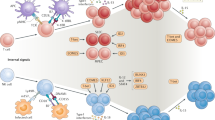
Key Points
Innate immunity has a long phylogenetic history, encompassing species as diverse as sea anemones, insects and mammals. By contrast, adaptive immunity, which involves lymphocyte-like cells and the antigen-binding receptors they express, is restricted to vertebrates.
A major departure in adaptive immunity is evident within vertebrate species. Jawed vertebrates use V(D)J recombination of immunoglobulin and T cell receptors, whereas jawless vertebrates rearrange variable lymphocyte receptors encoding leucine-rich repeats to form an alternative type of immune receptor.
Homologues of molecules that previously were thought to be unique to the rearrangement and diversification of immunoglobulin and T cell receptors have been identified in invertebrate species, in which these forms of immune recognition molecules are absent.
The enormously complex, multifactorial and highly regulated cellular processes that generate receptor variation in somatic cells arose through the integration of molecular systems that are not exclusively associated with immune diversity.
Reconstructing the nature of ancestral forms of adaptive immune receptors is compromised by the absence of crucial intermediates; however, it is possible to infer some of the main steps that gave rise to the antigen receptor-bearing immunocytes of jawed vertebrates.
Abstract
Adaptive immunity is mediated through numerous genetic and cellular processes that generate favourable somatic variants of antigen-binding receptors under evolutionary selection pressure by pathogens and other factors. Advances in our understanding of immunity in mammals and other model organisms are revealing the underlying basis and complexity of this remarkable system. Although the evolution of adaptive immunity has been thought to occur by the acquisition of novel molecular capabilities, an increasing amount of information from new model systems suggest that co-option and redirection of pre-existing systems are the main source of innovation. We combine evidence from a wide range of organisms to obtain an integrated view of the origins and patterns of divergence in adaptive immunity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Litman, G. W., Cannon, J. P. & Dishaw, L. J. Reconstructing immune phylogeny: new perspectives. Nature Rev. Immunol. 5, 866–879 (2005).
Flajnik, M. F. & Kasahara, M. Origin and evolution of the adaptive immune system: genetic events and selective pressures. Nature Rev. Genet. 11, 47–59 (2010).
Iwasaki, A. & Medzhitov, R. Regulation of adaptive immunity by the innate immune system. Science 327, 291–295 (2010).
Tonegawa, S. Somatic generation of antibody diversity. Nature 302, 575–581 (1983).
Gellert, M. V(D)J recombination: RAG proteins, repair factors, and regulation. Annu. Rev. Biochem. 71, 101–132 (2002).
Flajnik, M. F. Comparative analyses of immunoglobulin genes: surprises and portents. Nature Rev. Immunol. 2, 688–698 (2002).
Kokubu, F., Litman, R., Shamblott, M. J., Hinds, K. & Litman, G. W. Diverse organization of immunoglobulin VH gene loci in a primitive vertebrate. EMBO J. 7, 3413–3422 (1988).
Anderson, M. K., Shamblott, M. J., Litman, R. T. & Litman, G. W. Generation of immunoglobulin light chain gene diversity in Raja erinacea is not associated with somatic rearrangement, an exception to a central paradigm of B cell immunity. J. Exp. Med. 182, 109–119 (1995).
Ota, T., Rast, J. P., Litman, G. W. & Amemiya, C. T. Lineage-restricted retention of a primitive immunoglobulin heavy chain isotype within the Dipnoi reveals an evolutionary paradox. Proc. Natl Acad. Sci. USA 100, 2501–2506 (2003).
Zhao, Y. et al. Identification of IgF, a hinge-region-containing Ig class, and IgD in Xenopus tropicalis. Proc. Natl Acad. Sci. USA 103, 12087–12092 (2006).
Ohta, Y. & Flajnik, M. IgD, like IgM, is a primordial immunoglobulin class perpetuated in most jawed vertebrates. Proc. Natl Acad. Sci. USA 103, 10723–10728 (2006).
Richards, M. H. & Nelson, J. L. The evolution of vertebrate antigen receptors: a phylogenetic approach. Mol. Biol. Evol. 17, 146–155 (2000).
Rast, J. P. et al. α, β, γ, and δ T cell antigen receptor genes arose early in vertebrate phylogeny. Immunity 6, 1–11 (1997).
Miracle, A. L. et al. Complex expression patterns of lymphocyte-specific genes during the development of cartilaginous fish implicate unique lymphoid tissues in generating an immune repertoire. Int. Immunol. 13, 567–580 (2001).
Zapata, A. & Amemiya, C. T. Phylogeny of lower vertebrates and their immunological structures. Curr. Top. Microbiol. Immunol. 248, 67–107 (2000).
Pollara, B., Litman, G. W., Finstad, J., Howell, J. & Good, R. A. The evolution of the immune response. VII. Antibody to human “O” cells and properties of the immunoglobulin in lamprey. J. Immunol. 105, 738–745 (1970).
Litman, G. W., Anderson, M. K. & Rast, J. P. Evolution of antigen binding receptors. Ann. Rev. Immunol. 17, 109–147 (1999).
Alder, M. N. et al. Diversity and function of adaptive immune receptors in a jawless vertebrate. Science 310, 1970–1973 (2005).
Alder, M. N. et al. Antibody responses of variable lymphocyte receptors in the lamprey. Nature Immunol. 9, 319–327 (2008).
Saha, N. R., Smith, J. & Amemiya, C. T. Evolution of adaptive immune recognition in jawless vertebrates. Semin. Immunol. 22, 25–33 (2010).
Uinuk-Ool, T. et al. Lamprey lymphocyte-like cells express homologs of genes involved in immunologically relevant activities of mammalian lymphocytes. Proc. Natl Acad. Sci. USA 99, 14356–14361 (2002).
Mayer, W. E. et al. Isolation and characterization of lymphocyte-like cells from a lamprey. Proc. Natl Acad. Sci. USA 99, 14350–14355 (2002).
Bajoghli, B. et al. Evolution of genetic networks underlying the emergence of thymopoiesis in vertebrates. Cell 138, 186–197 (2009).
Pancer, Z. et al. Somatic diversification of variable lymphocyte receptors in the agnathan sea lamprey. Nature 430, 174–180 (2004). An important paper that reports the existence of a new diversified antigen receptor that is the basis for adaptive immunity in jawless vertebrates.
Pancer, Z. et al. Variable lymphocyte receptors in hagfish. Proc. Natl Acad. Sci. USA 102, 9224–9229 (2005).
Han, B. W., Herrin, B. R., Cooper, M. D. & Wilson, I. A. Antigen recognition by variable lymphocyte receptors. Science 321, 1834–1837 (2008).
Velikovsky, C. A. et al. Structure of a lamprey variable lymphocyte receptor in complex with a protein antigen. Nature Struct. Mol. Biol. 16, 725–730 (2009).
Smith, J. J., Antonacci, F., Eichler, E. E. & Amemiya, C. T. Programmed loss of millions of base pairs from a vertebrate genome. Proc. Natl Acad. Sci. USA 106, 11212–11217 (2009).
Nagawa, F. et al. Antigen-receptor genes of the agnathan lamprey are assembled by a process involving copy choice. Nature Immunol. 8, 206–213 (2007). The authors provide the first experimental data in support of a copy choice model for the assembly of VLR genes in jawless vertebrates.
Rogozin, I. B. et al. Evolution and diversification of lamprey antigen receptors: evidence for involvement of an AID–APOBEC family cytosine deaminase. Nature Immunol. 8, 647–656 (2007). The authors identify two cytidine deaminases with homology to AID and, based on the expression pattern, suggest that these enzymes are involved in the diversification of the VLR repertoire in lampreys.
Kishishita, N. et al. Regulation of antigen-receptor gene assembly in hagfish. EMBO Rep. 11, 126–132 (2010).
Tasumi, S. et al. High-affinity lamprey VLRA and VLRB monoclonal antibodies. Proc. Natl Acad. Sci. USA 106, 12891–12896 (2009).
Thompson, C. B. New insights into V(D)J recombination and its role in the evolution of the immune system. Immunity 3, 531–539 (1995).
Marchalonis, J. J., Schluter, S. F., Bernstein, R. M. & Hohman, V. S. Antibodies of sharks: revolution and evolution. Immunol. Rev. 166, 103–122 (1998).
Holland, L. Z. et al. The amphioxus genome illuminates vertebrate origins and cephalochordate biology. Genome Res. 18, 1100–1111 (2008).
Hibino, T. et al. The immune gene repertoire encoded in the purple sea urchin genome. Dev. Biol. 300, 349–365 (2006).
Azumi, K. et al. Genomic analysis of immunity in a Urochordate and the emergence of the vertebrate immune system: “waiting for Godot”. Immunogenetics 55, 570–581 (2003).
Eason, D. D., Litman, R. T., Luer, C. A., Kerr, W. & Litman, G. W. Expression of individual immunoglobulin genes occurs in an unusual system consisting of multiple independent loci. Eur. J. Immunol. 34, 2551–2558 (2004).
Fugmann, S. D., Messier, C., Novack, L. A., Cameron, R. A. & Rast, J. P. An ancient evolutionary origin of the Rag1/2 gene locus. Proc. Natl Acad. Sci. USA 103, 3728–3733 (2006). This is the first report of a RAG1 - and RAG2 -like gene cluster outside the jawed vertebrate lineage.
Kapitonov, V. V. & Jurka, J. RAG1 core and V(D)J recombination signal sequences were derived from transib transposons. PLoS Biol. 3, e181 (2005). This report identifies the striking similarity between RAG1 and Transib transposons, a widespread family of mobile DNA elements.
Fugmann, S. D. The origins of the Rag genes — from transposition to V(D)J recombination. Semin. Immunol. 22, 10–16 (2010).
Matthews, A. G. et al. RAG2 PHD finger couples histone H3 lysine 4 trimethylation with V(D)J recombination. Nature 450, 1106–1110 (2007).
Liu, Y., Subrahmanyam, R., Chakraborty, T., Sen, R. & Desiderio, S. A plant homeodomain in RAG-2 that binds hypermethylated lysine 4 of histone H3 is necessary for efficient antigen-receptor-gene rearrangement. Immunity 27, 561–571 (2007).
Wilson, D. R., Norton, D. D. & Fugmann, S. D. The PHD domain of the sea urchin RAG2 homolog, SpRAG2L, recognizes dimethylated lysine 4 in histone H3 tails. Dev. Comp. Immunol. 32, 1221–1230 (2008).
Ji, Y. et al. The in vivo pattern of binding of RAG1 and RAG2 to antigen receptor loci. Cell 141, 419–431 (2010).
Hinds-Frey, K. R., Nishikata, H., Litman, R. T. & Litman, G. W. Somatic variation precedes extensive diversification of germline sequences and combinatorial joining in the evolution of immunoglobulin heavy chain diversity. J. Exp. Med. 178, 825–834 (1993).
Diaz, M., Greenberg, A. S. & Flajnik, M. F. Somatic hypermutation of the new antigen receptor gene (NAR) in the nurse shark does not generate the repertoire: possible role in antigen-driven reactions in the absence of germinal centers. Proc. Natl Acad. Sci. USA 95, 14343–14348 (1998).
Lee, S. S., Tranchina, D., Ohta, Y., Flajnik, M. F. & Hsu, E. Hypermutation in shark immunoglobulin light chain genes results in contiguous substitutions. Immunity 16, 571–582 (2002).
Hayakawa, K. & Hardy, R. R. Development and function of B-1 cells. Curr. Opin. Immunol. 12, 346–353 (2000).
Takagaki, Y., Seipelt, R. L., Peterson, M. L. & Manley, J. L. The polyadenylation factor CstF-64 regulates alternative processing of IgM heavy chain pre-mRNA during B cell differentiation. Cell 87, 941–952 (1996).
Martincic, K., Alkan, S. A., Cheatle, A., Borghesi, L. & Milcarek, C. Transcription elongation factor ELL2 directs immunoglobulin secretion in plasma cells by stimulating altered RNA processing. Nature Immunol. 10, 1102–1109 (2009).
Sciammas, R. & Davis, M. M. Blimp-1; immunoglobulin secretion and the switch to plasma cells. Curr. Top. Microbiol. Immunol. 290, 201–224 (2005).
Sciammas, R. & Davis, M. M. Modular nature of Blimp-1 in the regulation of gene expression during B cell maturation. J. Immunol. 172, 5427–5440 (2004).
Heaney, M. L. & Golde, D. W. Soluble receptors in human disease. J. Leukoc. Biol. 64, 135–146 (1998).
Butler, J. E. Immunoglobulin diversity, B-cell and antibody repertoire development in large farm animals. Rev. Sci. Tech. 17, 43–70 (1998).
Longerich, S., Basu, U., Alt, F. & Storb, U. AID in somatic hypermutation and class switch recombination. Curr. Opin. Immunol. 18, 164–174 (2006).
Di Noia, J. M. & Neuberger, M. S. Molecular mechanisms of antibody somatic hypermutation. Annu. Rev. Biochem. 76, 1–22 (2007).
Dooley, H. & Flajnik, M. F. Shark immunity bites back: affinity maturation and memory response in the nurse shark, Ginglymostoma cirratum. Eur. J. Immunol. 35, 936–945 (2005).
Malecek, K. et al. Immunoglobulin heavy chain exclusion in the shark. PLoS Biol. 6, e157 (2008).
Harris, R. S. & Liddament, M. T. Retroviral restriction by APOBEC proteins. Nature Rev. Immunol. 4, 868–877 (2004).
Zarrin, A. A. et al. An evolutionarily conserved target motif for immunoglobulin class-switch recombination. Nature Immunol. 5, 1275–1281 (2004).
Weinstein, J. A., Jiang, N., White, R. A., Fisher, D. S. & Quake, S. R. Sequencing the zebrafish immune repertoire. Science 324, 807–810 (2009).
Chen, H. et al. Characterization of arrangement and expression of the T cell receptor γ locus in the sandbar shark. Proc. Natl Acad. Sci. USA 106, 8591–8596 (2009).
O'Brien, R. L. & Born, W. K. γδ T cell subsets: A link between TCR and function? Semin. Immunol. 5 May 2010 (doi:10.1016/j.smim.2010.03.006).
Van Rhijn, I. et al. Highly diverse TCR δ chain repertoire in bovine tissues due to the use of up to four D segments per δ chain. Mol. Immunol. 44, 3155–3161 (2007).
Boehm, T. Co-evolution of a primordial peptide-presentation system and cellular immunity. Nature Rev. Immunol. 6, 79–84 (2006).
Boehm, T. & Bleul, C. C. The evolutionary history of lymphoid organs. Nature Immunol. 8, 131–135 (2007).
Li, J. et al. B lymphocytes from early vertebrates have potent phagocytic and microbicidal abilities. Nature Immunol. 7, 1116–1124 (2006).
Guo, P. et al. Dual nature of the adaptive immune system in lampreys. Nature 459, 796–801 (2009). This work reports that VLR-expressing lymphocyte-like cells can be divided into two classes that are reminiscent of the humoral (B cells) and cellular (T cells) arms of the conventional adaptive immune system in jawed vertebrates.
Dermody, T. S., Kirchner, E., Guglielmi, K. M. & Stehle, T. Immunoglobulin superfamily virus receptors and the evolution of adaptive immunity. PLoS Pathog. 5, e1000481 (2009).
Yoder, J. A. et al. Resolution of the NITR gene cluster in zebrafish. Proc. Natl Acad. Sci. USA 101, 15706–15711 (2004).
Cannon, J. P. et al. A bony fish immunological receptor of the NITR multigene family mediates allogeneic recognition. Immunity 29, 228–237 (2008).
Cannon, J. P., Haire, R. N. & Litman, G. W. Identification of diversified genes that contain immunoglobulin-like variable regions in a protochordate. Nature Immunol. 3, 1200–1207 (2002). This article describes the identification of a highly diverse family of immunoglobulin receptors in amphioxus and raises the possibility that invertebrates may also rely on diversified antigen receptors for immunity, a characteristic previously thought to be exclusive to jawed vertebrates.
Cannon, J. P., Haire, R. N., Schnitker, N., Mueller, M. G. & Litman, G. W. Individual protochordates possess unique immune-type receptor repertoires. Curr. Biol. 14, R465–R466 (2004).
Hernandez Prada, J. A. et al. Ancient evolutionary origin of diversified variable regions revealed by crystal structures of an immune-type receptor in amphioxus. Nature Immunol. 7, 875–882 (2006).
Dishaw, L. J. et al. Genomic complexity of the variable region-containing chitin-binding proteins in amphioxus. BMC Genetics 9, 78 (2008).
Criscitiello, M. F., Saltis, M. & Flajnik, M. F. An evolutionarily mobile antigen receptor variable region gene: doubly rearranging NAR-TcR genes in sharks. Proc. Natl Acad. Sci. USA 103, 5036–5041 (2006).
Parra, Z. E. et al. A unique T cell receptor discovered in marsupials. Proc. Natl Acad. Sci. USA 104, 9776–9781 (2007).
Miller, R. D. Those other mammals: the immunoglobulins and T cell receptors of marsupials and monotremes. Semin. Immunol. 22, 3–9 (2010).
Rothenberg, E. V. & Pant, R. Origins of lymphocyte developmental programs: transcription factor evidence. Semin. Immunol. 16, 227–238 (2004).
Anderson, S. K. Transcriptional regulation of NK cell receptors. Curr. Top. Microbiol. Immunol. 298, 59–75 (2006).
Rast, J. P., Smith, L. C., Loza-Coll, M., Hibino, T. & Litman, G. W. Genomic insights into the immune system of the sea urchin. Science 314, 952–956 (2006).
Pancer, Z. & Cooper, M. D. The evolution of adaptive immunity. Ann. Rev. Immunol. 24, 497–518 (2006).
Zhang, Q. et al. Novel genes dramatically alter regulatory network topology in amphioxus. Genome Biol. 9, R123 (2008).
Hedrick, S. M. The acquired immune system: a vantage from beneath. Immunity 21, 607–615 (2004).
Janeway, C. A. Jr & Medzhitov, R. Innate immune recognition. Annu. Rev. Immunol. 20, 197–216 (2002).
Huang, S. et al. Genomic analysis of the immune gene repertoire of amphioxus reveals extraordinary innate complexity and diversity. Genome Res. 18, 1112–1126 (2008).
Davidson, C. R., Best, N. M., Francis, J. W., Cooper, E. L. & Wood, T. C. Toll-like receptor genes (TLRs) from Capitella capitata and Helobdella robusta (Annelida). Dev. Comp. Immunol. 32, 608–612 (2008).
Messier-Solek, C., Buckley, K. M. & Rast, J. P. Highly diversified innate receptor systems and new forms of animal immunity. Semin. Immunol. 22, 39–47 (2010).
Flajnik, M. F. & Du, P. L. Evolution of innate and adaptive immunity: can we draw a line? Trends Immunol. 25, 640–644 (2004).
Lieber, M. R. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu. Rev. Biochem. 79, 181–211 (2010).
Weller, G. R. et al. Identification of a DNA nonhomologous end-joining complex in bacteria. Science 297, 1686–1689 (2002).
Meek, K., Gupta, S., Ramsden, D. A. & Lees-Miller, S. P. The DNA-dependent protein kinase: the director at the end. Immunol. Rev. 200, 132–141 (2004).
Gozalbo-Lopez, B. et al. A role for DNA polymerase μ in the emerging DJH rearrangements of the postgastrulation mouse embryo. Mol. Cell Biol. 29, 1266–1275 (2009).
Davidson, E. H. & Erwin, D. H. Gene regulatory networks and the evolution of animal body plans. Science 311, 796–800 (2006).
Acknowledgements
The work in the laboratory of G.W.L. is supported by US National Institutes of Health grants AI23338 and AI57559. The work in the laboratory of J.P.R. is supported by Canadian Institutes for Health Research grant MOP74667 and National Science and Engineering Research Council of Canada grant NSERC 458115/211598. The work in the laboratory of S.D.F. is supported by the Intramural Research Program of the US National Institutes of Health, National Institute on Aging.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- Somatic hypermutation
-
(SHM). The process by which point mutations are introduced in the heavy- or light-chain variable region gene segments, resulting in a change in the expressed protein, which may alter affinity or specificity for antigen.
- V(D)J recombination
-
The lineage-specific RAG1- and RAG2-mediated assembly of functional immunoglobulin and T cell receptor genes from individual variable (V), diversity (D) and joining (J) gene segments. This process occurs exclusively in B and T cell progenitors.
- Recombination signal sequences
-
(RSSs). Highly conserved heptamer and nonomer sequences that flank recombining segmental elements in immunoglobulin and T cell receptor genes. These sequences are separated by 12 or 23 base pair spacers that are required to direct the recombination of compatible elements.
- Recombination-activating gene 1
-
(RAG1). A protein that, along with RAG2, forms a DNA recombinase complex which catalyses V(D)J recombination. During the joining phase additional ubiquitous DNA repair factors are recruited. Both genes are exclusively expressed in developing lymphocytes.
- Gene conversion
-
An immunoglobulin gene diversification process closely related to somatic hypermutation. In general, U:G base mismatches are repaired by copying sequence information from upstream pseudo V segments into the rearranged VJ and VDJ exon of the IgL and IgH genes, respectively. In birds and rabbits it occurs before antigen encounter.
- Tetrapods
-
Four-limbed vertebrates that include the amphibians and amniotes (reptiles, birds and mammals). These groups share some common immune features, such as class-switch recombination.
- Activation-induced cytidine deaminase
-
(AID). An enzyme that acts on single-stranded DNA and converts cytidines to uracils. It is typically expressed in B cells after antigen encounter. Its mutagenic activity is restricted to immunoglobulin loci.
- Phylogeny
-
The study of evolutionary relatedness among groups or species of organisms. Similar analyses can be applied to relatedness among genes.
- Lampreys and hagfish
-
Two extant groups of the jawless vertebrate lineage that have anatomical and physiological differences from jawed vertebrates. Lampreys undergo metamorphosis from a larval form, termed an ammocoete, and approximately half of the species are parasitic in adult life. Hagfish are ocean dwelling species that feed on decaying animal matter and small crustaceans.
- Leucine-rich repeat
-
(LRR). A protein structural motif that is comprised of 20–30 amino acid regions that are unusually rich in the hydrophobic amino acid leucine and form a characteristic structural fold.
- Copy choice mechanism of recombination
-
A genetic recombination mechanism in which a new DNA sequence is generated by replication using multiple DNA sequences as templates. The DNA polymerase continues to synthesize a single DNA molecule while jumping from one template strand to another. The choice of templates is largely driven by sequence homologies, allowing the previously synthesized DNA to anneal to similar sequences elsewhere in the genome and thereby redirecting DNA synthesis.
- Chordates
-
An animal phylum that comprises both jawed and jawless vertebrates, and two subphyla of invertebrates: the cephalochordates (such as amphioxus) and the urochordates (such as the sea squirt Ciona intestinalis).
- Deuterostomes
-
An animal superphylum composed of four phyla: the chordates (which include vertebrates), the echinoderms (consisting of starfish, sea urchins and allied species), the hemichordates (acorn worms) and Xenoturbellida (containing two marine worm-like species). Genomes from each of the three main deuterostome phyla have been sequenced.
- Transposons
-
(Also are known as 'jumping genes' and 'selfish DNA'). DNA sequences that encode transposases, the enzymes required to excise the transposon from its original chromosomal location and to integrate it in a different position within the genome. The ends of transposons consist of DNA repeats that serve as recognition sites for the transposase itself.
- Purple sea urchin
-
(Strongylocentrotus purpuratus). An echinoderm that is a well established model system for developmental biology. Sequencing of its genome increased the interest of the broader scientific community (including immunologists) in this invertebrate model system.
- Complementarity determining region 3
-
(CDR3). The region in B and T cell receptors that interacts with the antigen, in which hypervariable sequences are located. CDR3 typically is derived somatically, whereas CDR1 and CDR2 are germline encoded.
- B-1 B cells
-
In mice, this B cell subset is found mainly in the peritoneal and pleural cavities. B-1 B cells are a self-renewing subset with a restricted repertoire of B cell receptors that respond to common bacterial antigens and have a role in autoimmunity.
- Derived evolutionary features
-
Anatomical structures, genes or functional systems are designated as derived when they originate within a sub-lineage. Primitive evolutionary features refer to those that originate from similar features in a common ancestor.
- Invariant natural killer T (iNKT) cells
-
Lymphocytes that express a particular variable gene segment, Vα14 (in mice) and Vα24 (in humans), precisely rearranged to a particular Jα gene segment to yield T cell receptor α-chains with an invariant sequence.
- Novel immune-type receptors
-
(NITRs). Activating or inhibitory receptors that are encoded by large diversified multigene families in bony fish. All NITRs have a variable region and most have a transmembrane region and cytoplasmic tail with signalling motifs.
Rights and permissions
About this article
Cite this article
Litman, G., Rast, J. & Fugmann, S. The origins of vertebrate adaptive immunity. Nat Rev Immunol 10, 543–553 (2010). https://doi.org/10.1038/nri2807
Issue Date:
DOI: https://doi.org/10.1038/nri2807
This article is cited by
Origin and evolutionary malleability of T cell receptor α diversity
Nature (2023)
Ancient fish lineages illuminate toll-like receptor diversification in early vertebrate evolution
Immunogenetics (2023)
On the relationship between extant innate immune receptors and the evolutionary origins of jawed vertebrate adaptive immunity
Immunogenetics (2022)
Immunoglobulin-like receptors and the generation of innate immune memory
Immunogenetics (2022)
Regional effect on the molecular clock rate of protein evolution in Eutherian and Metatherian genomes
BMC Ecology and Evolution (2021)