Abstract
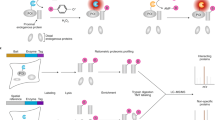
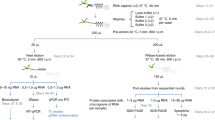
RNA–protein interactions play numerous roles in cellular function and disease. Here we describe RNA–protein interaction detection (RaPID), which uses proximity-dependent protein labeling, based on the BirA* biotin ligase, to rapidly identify the proteins that bind RNA sequences of interest in living cells. RaPID displays utility in multiple applications, including in evaluating protein binding to mutant RNA motifs in human genetic disorders, in uncovering potential post-transcriptional networks in breast cancer, and in discovering essential host proteins that interact with Zika virus RNA. To improve the BirA*-labeling component of RaPID, moreover, a new mutant BirA* was engineered from Bacillus subtilis, termed BASU, that enables >1,000-fold faster kinetics and >30-fold increased signal-to-noise ratio over the prior standard Escherichia coli BirA*, thereby enabling direct study of RNA–protein interactions in living cells on a timescale as short as 1 min.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
20 April 2018
In the version of this Article originally published, an amino acid substitution introduced into the B. subtilis biotin ligase sequence was incorrectly written as R142G. The correct mutation is R124G. This error has been corrected in the HTML and PDF versions of the paper.
References
Moore, M.J. From birth to death: the complex lives of eukaryotic mRNAs. Science 309, 1514–1518 (2005).
Keene, J.D. RNA regulons: coordination of post-transcriptional events. Nat. Rev. Genet. 8, 533–543 (2007).
Castello, A. et al. Insights into RNA biology from an atlas of mammalian mRNA-binding proteins. Cell 149, 1393–1406 (2012).
Lee, J.T. Epigenetic regulation by long noncoding RNAs. Science 338, 1435–1439 (2012).
Wang, P. et al. The STAT3-binding long noncoding RNA lnc-DC controls human dendritic cell differentiation. Science 344, 310–313 (2014).
Keefe, A.D., Pai, S. & Ellington, A. Aptamers as therapeutics. Nat. Rev. Drug Discov. 9, 537–550 (2010).
Galloway, K.E., Franco, E. & Smolke, C.D. Dynamically reshaping signaling networks to program cell fate via genetic controllers. Science 341, 1235005 (2013).
Butter, F., Scheibe, M., Mörl, M. & Mann, M. Unbiased RNA-protein interaction screen by quantitative proteomics. Proc. Natl. Acad. Sci. USA 106, 10626–10631 (2009).
Mili, S. & Steitz, J.A. Evidence for reassociation of RNA-binding proteins after cell lysis: implications for the interpretation of immunoprecipitation analyses. RNA 10, 1692–1694 (2004).
Matia-González, A.M., Iadevaia, V. & Gerber, A.P. A versatile tandem RNA isolation procedure to capture in vivo formed mRNA-protein complexes. Methods 118-119, 93–100 (2017).
Rogell, B. et al. Specific RNP capture with antisense LNA/DNA mixmers. RNA 23, 1290–1302 (2017).
McHugh, C.A. et al. The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3. Nature 521, 232–236 (2015).
Buenrostro, J.D. et al. Quantitative analysis of RNA-protein interactions on a massively parallel array reveals biophysical and evolutionary landscapes. Nat. Biotechnol. 32, 562–568 (2014).
Kim, D.I. et al. Probing nuclear pore complex architecture with proximity-dependent biotinylation. Proc. Natl. Acad. Sci. USA 111, E2453–E2461 (2014).
Roux, K.J., Kim, D.I., Raida, M. & Burke, B. A promiscuous biotin ligase fusion protein identifies proximal and interacting proteins in mammalian cells. J. Cell Biol. 196, 801–810 (2012).
Austin, R.J., Xia, T., Ren, J., Takahashi, T.T. & Roberts, R.W. Designed arginine-rich RNA-binding peptides with picomolar affinity. J. Am. Chem. Soc. 124, 10966–10967 (2002).
Edwards, J.M., Long, J., de Moor, C.H., Emsley, J. & Searle, M.S. Structural insights into the targeting of mRNA GU-rich elements by the three RRMs of CELF1. Nucleic Acids Res. 41, 7153–7166 (2013).
Zheng, X. et al. Detecting RNA–protein interaction using end-labeled biotinylated RNA oligonucleotides and immunoblotting 1421, 35–44 (2016).
Mellacheruvu, D. et al. The CRAPome: a contaminant repository for affinity purification-mass spectrometry data. Nat. Methods 10, 730–736 (2013).
Allerson, C.R., Cazzola, M. & Rouault, T.A. Clinical severity and thermodynamic effects of iron-responsive element mutations in hereditary hyperferritinemia-cataract syndrome. J. Biol. Chem. 274, 26439–26447 (1999).
Wang, J. et al. Iron-dependent degradation of apo-IRP1 by the ubiquitin-proteasome pathway. Mol. Cell. Biol. 27, 2423–2430 (2007).
Mlakar, J. et al. Zika virus associated with microcephaly. N. Engl. J. Med. 374, 951–958 (2016).
Li, Z. & Nagy, P.D. Diverse roles of host RNA binding proteins in RNA virus replication. RNA Biol. 8, 305–315 (2011).
Tang, H. et al. Zika virus infects human cortical neural progenitors and attenuates their growth. Cell Stem Cell 18, 587–590 (2016).
Uhlén, M. et al. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Hardy, R.J. QKI expression is regulated during neuron-glial cell fate decisions. J. Neurosci. Res. 54, 46–57 (1998).
Höistad, M. et al. Linking white and grey matter in schizophrenia: oligodendrocyte and neuron pathology in the prefrontal cortex. Front. Neuroanat. 3, 9 (2009).
Parker, B.J. et al. New families of human regulatory RNA structures identified by comparative analysis of vertebrate genomes. Genome Res. 21, 1929–1943 (2011).
Oikonomou, P., Goodarzi, H. & Tavazoie, S. Systematic identification of regulatory elements in conserved 3′ UTRs of human transcripts. Cell Rep. 7, 281–292 (2014).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013).
Pratama, A. et al. Roquin-2 shares functions with its paralog Roquin-1 in the repression of mRNAs controlling T follicular helper cells and systemic inflammation. Immunity 38, 669–680 (2013).
Janowski, R. et al. Roquin recognizes a non-canonical hexaloop structure in the 3′-UTR of Ox40. Nat. Commun. 7, 11032 (2016).
Kim, D.I. et al. An improved smaller biotin ligase for BioID proximity labeling. Mol. Biol. Cell 27, 1188–1196 (2016).
Henke, S.K. & Cronan, J.E. Successful conversion of the Bacillus subtilis BirA Group II biotin protein ligase into a Group I ligase. PLoS One 9, e96757 (2014).
Schlundt, A. et al. Structural basis for RNA recognition in roquin-mediated post-transcriptional gene regulation. Nat. Struct. Mol. Biol. 21, 671–678 (2014).
Lam, S.S. et al. Directed evolution of APEX2 for electron microscopy and proximity labeling. Nat. Methods 12, 51–54 (2015).
Zhang, J.-P. et al. Efficient precise knockin with a double cut HDR donor after CRISPR/Cas9-mediated double-stranded DNA cleavage. Genome Biol. 18, 35 (2017).
Ramanathan, M. & Khavari, P.A. RNA-protein interaction detection. Protocol Exchange https://doi.org/10.1038/protex.2018.003 (2018).
Rappsilber, J., Ishihama, Y. & Mann, M. Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal. Chem. 75, 663–670 (2003).
Okita, K. et al. A more efficient method to generate integration-free human iPS cells. Nat. Methods 8, 409–412 (2011).
Shi, Y., Kirwan, P., Smith, J., Robinson, H.P.C. & Livesey, F.J. Human cerebral cortex development from pluripotent stem cells to functional excitatory synapses. Nat. Neurosci. 15, 477–486 (2012).
Sharma, A. et al. Human induced pluripotent stem cell-derived cardiomyocytes as an in vitro model for coxsackievirus B3-induced myocarditis and antiviral drug screening platform. Circ. Res. 115, 556–566 (2014).
Zarnegar, B.J. et al. irCLIP platform for efficient characterization of protein-RNA interactions. Nat. Methods 13, 489–492 (2016).
Ridky, T.W., Chow, J.M., Wong, D.J. & Khavari, P.A. Invasive three-dimensional organotypic neoplasia from multiple normal human epithelia. Nat. Med. 16, 1450–1455 (2010).
Acknowledgements
We thank C. Adams, R. Leib and Vincent Coates Foundation Mass Spectrometry Laboratory, Stanford University Mass Spectrometry for help with mass spectrometry. We also thank P. Oikonomou and S. Tavazoie for providing position weight matrixes. We thank A. Fire, H. Chang, J. Elias, P. Sarnow, R. Flynn, J. Quinn, A. Bhaduri, A. Rubin and M. Kay for presubmission review. We thank L. Morcom and P. Bernstein for expert administrative assistance and members of the Khavari lab for helpful discussions. This work was supported by the US VA Office of Research and Development, by NIH AR49737, and NIH AR43799 (P.A.K.). The project described was supported by Award Number S10RR027425 from the National Center For Research Resources. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Center for Research Resources or the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
M.R. designed and executed experiments, analyzed the data and wrote the manuscript. K.M. designed and performed viral replication assays. J.G.R., H.G., D.S.R., J.R.K., P.H.N., B.J.Z., Z.S. and S.M. performed experiments. J.E.C. and T.D.P. supervised work. P.A.K. designed experiments, analyzed data, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Validation of RaPID with EDEN15
(a) Full Western Blot of conventional biotinylated RNA pulldown with EDEN15 and scrambled control (n=3 biologically independent experiments, representative image shown) (b) Schema of motif plasmid. Motif along with BoxB sites added to 3′UTR of eGFP (c) Quantification of motif plasmid expression in HEK293T cells by qPCR with primers against eGFP. (n=3 biologically independent experiments, bars represent mean signal and error bars denote s.e.m) (d) RaPID-Western of EDEN15 vs two scrambled sequence controls of identical nucleotide composition (scr1 and scr2). HA recognizes the epitope tagged LN-HA-BirA* fusion, which serves as a self-biotinylated streptavidin pulldown internal control (n=3 biologically independent experiments, representative image shown)
Supplementary Figure 2 RaPID-Western IRE-IREB2 binding with HHCS mutants
(a) RaPID-Western blots of WT IRE and HHCS FTL IRE mutants. Western blotting of IREB2 pulldown, RaPID (λN-HA-BirA*) fusion protein, which serves as an self-biotinylated internal control detected by HA antibody, and IREB2 in the lysate are shown. (n=3 biologically independent experiments, representative image shown) (b) Quantification of motif plasmid expression in HEK293T cells by qPCR with primers against eGFP. (n=3 biologically independent experiments, bars represent mean signal and error bars denote s.e.m)
Supplementary Figure 3 RaPID detects disruption in IRE-IRP1 interaction upon drug treatment
RaPID-Western of IRE motif with treatment (deferoxamine [DFO] 100uM, dimethyl sulfoxide [DMSO] 0.1%, ferric ammonium citrate [FAC] 100uM). (a) RaPID-Western with IRE blotting for IRP1 with different drug treatments. (n=3 biologically independent experiments, representative image shown) (b) Quantification of RaPID-Westerns with drug treatments. (n=3 biologically independent experiments, bars represent mean signal and error bars denote s.e.m) (c) Full Western blot of representative image shown
Supplementary Figure 4 Analysis of RaPID-MS ZIKV UTR bound proteins
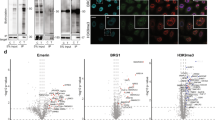
(a) GO Analysis performed on downregulated genes in ZIKV infected NPCs. RNA-Seq data used for analysis obtained from Tang et al.24 (b) Tissue enrichment score plot (please see methods) of RaPID-MS ZIKV UTR bound proteins. (c) Western Blot of U87 cells comparing efficacy of knockdown of QKI using scrambled with QKI shRNA 1 and QKI shRNA 2. (n=3 biologically independent experiments, representative image shown) (d) Quantification of QKI knockdown in U87 cells with Ctrl shRNA, QKI shRNA 1 and QKI shrNA 2. (n=3 biologically independent experiments, bars represent mean signal and error bars denote s.e.m) (e) Full Western blot of representative image shown
Supplementary Figure 5 QKI Protein levels decrease as NPCs differentiate to Neurons
(a) hiPSC colonies exhibit the pluripotency marker Oct4 (Green). (b) Day 21 NPCs are organized in a radial arrangement around the adherens junction marker NCad (Red) and are positive for the neuroectodermal marker Pax6 (Green) (c) Day 45 neurons are positive for the neuronal markers Tuj1 (Red) and NeuN (Green) (d) Western Blot of QKI comparing levels in hiPSC derived NPCs and neurons. (n=3 biologically independent experiments, representative image shown) (e) Quantification of QKI levels in hiPSC derived NPCs and neurons. (n=3 biologically independent experiments, bars represent mean signal and error bars denote s.e.m) (f) Full Western blot of representative image shown
Supplementary Figure 7 RC3H1&2 upergulation is correlated with poor survival outcome in breast cancer
RC3H1&2 upregulation correlated with poor survival outcomes in breast cancer (a) Breast cancer patient survival as a function of RC3H1 upregulation (red line) versus control (blue line). TCGA data - Breast Invasive Carcinoma (TCGA, Provisional) accessed on 5/2/2016 via cBioPortal with query settings of copy number alterations and mRNA expression (z-score>3). Logrank Test P-value calculated by cbioportal noted on plot (b) Breast cancer patient survival as a function of RC3H2 upregulation (red line) versus control (blue line). TCGA data - Breast Invasive Carcinoma (TCGA, Provisional) accessed on 5/2/2016 via cBioPortal with query settings of copy number alterations and mRNA expression (z-score>3).Logrank Test P-value calculated by cbioportal noted on plot (c) Breast cancer patient survival as a function of RC3H1 and RC3H2 upregulation (red line) versus control (blue line). TCGA data - Breast Invasive Carcinoma (TCGA, Provisional) accessed on 5/2/2016 via cBioPortal with query settings of copy number alterations and mRNA expression (z-score>3). Upregulation of RC3H1 and RC3H2 is mutually exclusive (Log Odds Ratio <-3 & Fisher Exact test p-value 0.035). Logrank Test P-value calculated by cbioportal noted on plot
Supplementary Figure 8 Structure and Sequences of Biotin ligases
Structure and sequences of biotin ligases (a) E. coli BirA (PDB:1BIB) annotated with color-coded motifs (RBAM, ABM, AM & PTAM) using MacPyMol (b) Sequences of RBAM, ABM, AM & PTAM in E. coli BirA (c) Sequence alignment of screened biotin ligases (A. aeolicus, Rhodanobacter spp., Tanerella spp., T. brucei) with motifs (RBAM, ABM, AM & PTAM) (d) Sequence alignment of N terminal truncated B. subtilis biotin ligase (67-325) with motifs (RBAM, ABM, AM & PTAM) (e) Alignment of B. subtilis biotin ligase and E. coli BirA*. Mutations made at C-terminus of B. subtilis (67-325) highlighted in red boxes to create BASU with SLR terminal end similar to E. coli BirA*.
Supplementary Figure 9 Expression and Streptavidin signal of HA-tagged mutant BirA*s
(A) HA signal of total cell lysate with each mutant biotin ligase (lower boxed area) transfected in HEK293T cells. Vinculin is blotted (upper boxed area) and used as loading control. (n=3 biologically independent experiments, representative image shown) (B) Streptavidin signal of total cell lysate with each mutant biotin ligase (n=3 biologically independent experiments, representative image shown)
Supplementary Figure 10 Evaluating application of BASU in RaPID with varying labeling timss
(a) RaPID-Western blots of TNF-CDE at different labeling time points ranging from 0min (unlabeled) to 1hour labeling. Western blotting of RC3H1 pulldown, RaPID (λN-HA-BirA*) fusion protein, which serves as an self-biotinylated internal control detected by HA antibody, and RC3H1 in the lysate are shown. (n=3 biologically independent experiments, representative image shown) (b) Full Western blot of representative image shown
Supplementary Figure 11 Higher signal-to-noise ratio with BASU shorter labeling
(a) Quantification of motif plasmid expression in HEK293T cells by qPCR with primers against eGFP. (n=3 biologically independent experiments, bars represent mean signal and error bars denote s.e.m) (b) RaPID-Western of TNF-CDE vs two scrambled sequence controls (scr1 and scr2). HA recognizes the epitope tagged LN-HA-BioID2* fusion, which serves as a self-biotinylated streptavidin pulldown internal control. Labeling performed for 1 min (n=3 biologically independent experiments, representative image shown) (c) RaPID-Western of TNF-CDE vs two scrambled sequence controls (scr1 and scr2). HA recognizes the epitope tagged LN-HA-BASU* fusion, which serves as a self-biotinylated streptavidin pulldown internal control. Labeling performed for 1 min (n=3 biologically independent experiments, representative image shown) (d) Quantification of RaPID-Westerns with BASU 1 min labeling. (n=3 biologically independent experiments, bars represent mean signal and error bars denote s.e.m) (e) RaPID-Western blots of TNF-CDE and scrambled controls using conventional E. coli BirA* with 18hrs of labeling. Western blotting of RC3H1 pulldown, RaPID (λN-HA-BirA*) fusion protein, which serves as an self-biotinylated internal control detected by HA antibody, and RC3H1 in the lysate are shown. (n=3 biologically independent experiments, representative image shown) (f) Quantification of RaPID-Western of TNF-CDE and scrambled controls using conventional E. coli BirA* with 18hrs of labeling (n=3 biologically independent experiments, bars represent mean signal and error bars denote s.e.m) (g) Signal-to-noise ratio (SNR) comparison between E. coli BirA* at 18hrs and BASU at 1min using TNF-CDE vs scrambled control RNA motifs (h) RaPID-MS SAINT score vs Fold Change (FC) plot for EDEN15 with BASU at labeling time of 30 minutes (i) RaPID-MS SAINT score vs Fold Change (FC) plot for IRE (UTRP 35) with BASU at labeling time of 30 minutes.
Supplementary Figure 12 Characterization and comparison of E.coli BirA* and BASU
(a) Immunoflourescence of streptavidin in 3D organotypic human skin tissue expressing BASU or E.coli BirA* taken at timepoint 0 min (no biotin labeling) and 30mins (30mins of biotin labeling). (b) Western Blot of streptavidin in 3D organotypic human skin tissue expressing BASU or E.coli BirA* taken at timepoint 0 min (no biotin labeling) and 30mins (30mins of biotin labeling). (n=3 biologically independent experiments, representative image shown) (c) Comparison of gain in streptavidin signal between E.coli BirA* and BASU. Normalized Streptavidin levels (Streptavidin level in Western Blot/ HA levels in Western Blot) were obtained for E.coli BirA* and BASU at each timepoint. The gain in streptavidin signal was obtained by subtracting Normalized Streptavidin levels at timepoint 0 min from timepoint 30mins for E.coli BirA* and BASU.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12 and Supplementary Notes 1–2 (PDF 3022 kb)
Supplementary Protocol
RaPID-Western (PDF 214 kb)
Supplementary Table 2
EDEN15 RaPID-MS in HEK293T cells. (XLSX 216 kb)
Supplementary Table 3
EDEN15 RaPID-MS in Huh7 cells. (XLSX 102 kb)
Supplementary Table 4
FTL IRE RaPID-MS in HEK293T cells. (XLSX 298 kb)
Supplementary Table 5
ZIKV 5′UTR RaPID-MS in HEK293T cells. (XLSX 271 kb)
Supplementary Table 6
ZIKV 3′UTR RaPID-MS in HEK293T cells. (XLSX 255 kb)
Supplementary Table 7
ZIKV UTR host proteins DAVID GO Term Analysis. (XLSX 83 kb)
Supplementary Table 8
Tissue Protein Level data for ZIKV UTR interacting proteins. (XLSX 47 kb)
Supplementary Table 9
Histone Stem Loop (UTRP11) RaPID-MS in HEK293T cells. (XLSX 214 kb)
Supplementary Table 10
p38_NRE RaPID-MS in HEK293T cells. (XLSX 179 kb)
Supplementary Table 11
PPP1R3C (UTRP30) RaPID-MS in HEK293T cells. (XLSX 220 kb)
Supplementary Table 12
Synthetic-EIR concatamer RaPID-MS in HEK293T cells. (XLSX 204 kb)
Supplementary Table 13
SM1v1 RaPID-MS in HEK293T cells. (XLSX 218 kb)
Supplementary Table 14
BASU EDEN15 RaPID-MS in HEK293T cells at 30 minutes. (XLSX 184 kb)
Supplementary Table 15
FTL IRE RaPID-MS in HEK293T cells at 30 minutes. (XLSX 208 kb)
Supplementary Table 16
List of Background proteins in HEK293T cells (XLSX 74 kb)
Supplementary Table 17
RNA motif sequences utilized in RaPID. (XLSX 58 kb)
Supplementary Table 18
Composition of buffers used in RaPID. (XLSX 41 kb)
Source data
Rights and permissions
About this article
Cite this article
Ramanathan, M., Majzoub, K., Rao, D. et al. RNA–protein interaction detection in living cells. Nat Methods 15, 207–212 (2018). https://doi.org/10.1038/nmeth.4601
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4601
This article is cited by
TREX reveals proteins that bind to specific RNA regions in living cells
Nature Methods (2024)
Profiling the interactome of oligonucleotide drugs by proximity biotinylation
Nature Chemical Biology (2024)
The rapid proximity labeling system PhastID identifies ATP6AP1 as an unconventional GEF for Rheb
Cell Research (2024)
TurboID screening of ApxI toxin interactants identifies host proteins involved in Actinobacillus pleuropneumoniae-induced apoptosis of immortalized porcine alveolar macrophages
Veterinary Research (2023)
The development of proximity labeling technology and its applications in mammals, plants, and microorganisms
Cell Communication and Signaling (2023)