-
PDF
- Split View
-
Views
-
Cite
Cite
Eun Mi Jung, Jun Hee Lim, Tae Jin Lee, Jong-Wook Park, Kyeong Sook Choi, Taeg Kyu Kwon, Curcumin sensitizes tumor necrosis factor-related apoptosis-inducing ligand (TRAIL)-induced apoptosis through reactive oxygen species-mediated upregulation of death receptor 5 (DR5), Carcinogenesis, Volume 26, Issue 11, November 2005, Pages 1905–1913, https://doi.org/10.1093/carcin/bgi167
Close - Share Icon Share
Abstract
Curcumin exhibits anti-inflammatory and antitumor activities. Although its functional mechanism has not been elucidated so far, numerous studies have shown that curcumin induces apoptosis in cancer cells. In the present study, we show that subtoxic concentrations of curcumin sensitize human renal cancer cells to the tumor necrosis factor-related apoptosis inducing ligand (TRAIL)-mediated apoptosis. This apoptosis induced by the combination of curcumin and TRAIL is not interrupted by Bcl-2 overexpression. We found that treatment with curcumin significantly induces death receptor 5 (DR5) expression both at its mRNA and protein levels, accompanying the generation of the reactive oxygen species (ROS). Not only the pretreatment with N -acetylcystine but also the ectopic expression of peroxiredoxin II, an antioxidative protein, dramatically inhibited the apoptosis induced by curcumin and TRAIL in combination, blocking the curcumin-mediated DR5 upregulation. Taken together, the present study demonstrates that curcumin enhances TRAIL-induced apoptosis by ROS-mediated DR5 upregulation.
Introduction
Curcumin is a major component of the Curcuma species, which is commonly used as a yellow coloring and flavoring agent in foods. Recently, several studies have shown that curcumin possesses antiproliferative activity against tumor cells in vitro ( 1 ). Moreover, curcumin exhibits anticarcinogenic activity in vivo , as indicated by its inhibition of tumor initiation induced by various carcinogens or phorbol esters ( 2 – 4 ). Although curcumin-induced cancer chemopreventive activity have been repeatedly reported in various cancer cell lines including skin, oral, intestinal, breast and colon cancer cells ( 5 – 8 ), the mechanisms underlying the diverse effects of curcumin on cancer cells are not fully understood.
Recently, it has been reported that curcumin treatment results in the production of reactive oxygen species (ROS) ( 9 , 10 ). ROS is involved in the initiation and progression of a variety of human diseases ( 11 ), including renal ischemia/reperfusion injury and in toxicities associated with chemical exposure. The kidney or kidney-derived cells have an increased susceptibility towards agents generating ROS in terms of apoptogenecity and proliferation, and ROS play an important role in the pathogenesis of a variety of renal diseases ( 12 ). Renal cell carcinoma remains one of the most drug-resistant malignancies in humans and is a frequent cause of cancer mortality ( 13 ), necessitating the development of novel therapeutic strategies against this type of cancer. In this study, we investigated the relationship between the curcumin-induced oxidative response and toxicity in renal carcinoma cells, the human renal Caki cells.
Tumor necrosis factor (TNF) α-related apoptosis-inducing ligand (TRAIL), also known as Apo2L, is a member of the TNF family and can induce apoptotic cell death in a variety of cancer cell types ( 14 – 16 ). TRAIL binds to four different types of membrane-bound death receptors (DR4, DR5, DcR1 and DcR2). Both DR4 and DR5 contain a conserved cytoplasmic region called the ‘death domain’, which is required for TRAIL-induced apoptosis ( 17 – 19 ). However, DcR1 lacks an intracellular domain, and DcR2 contains a truncated death domain. Thus, DcR1 and DcR2 may protect cells from TRAIL-induced apoptosis by acting as decoy receptors ( 18 , 20 ). While TRAIL induces apoptosis in a wide variety of tumor cells, it does not cause toxicity to most normal cells. This is because of the presence of large numbers of decoy receptors on normal cells ( 18 , 20 ). Recent studies have shown that some cancer cells are resistant to the apoptotic effects of TRAIL ( 21 – 23 ). However, TRAIL-resistant cancer cells can be sensitized by the combined treatment with chemotherapeutic drugs and TRAIL in vitro , indicating that combination therapy may overcome TRAIL-resistance in cancer cells. Therefore, both the understanding of the molecular mechanisms of TRAIL resistance and development of the strategies to sensitize these cancer cells to undergo TRAIL-mediated apoptosis are important issues for effective cancer therapy.
In this study, we investigated whether curcumin can enhance TRAIL-mediated apoptosis in human renal cancer cells. Curcumin treatment resulted in the upregulation of DR5 accompanying ROS generation and rendered cells more sensitive to the cytotoxic activity of TRAIL. Pretreatment with antioxidant N -acetylcystine (NAC) and ectopic expression of an antioxidative protein [peroxiredoxin II (Prx II)] markedly prevents curcumin-induced DR5 upregulation and apoptosis induced by subtoxic concentrations of curcumin and TRAIL in combination. Therefore, curcumin-induced ROS generation may be critical for DR5 upregulation and the subsequent activation of TRAIL-mediated apoptotic signaling pathways.
Materials and methods
Cells and materials
Caki, HCT116, HT29, Hep G2 and Hep 3B cells were obtained from the American Type Culture Collection (ATCC: Rockville, MD). The culture medium used throughout these experiments was Dulbecco's modified Eagle's medium (DMEM), containing 10% fetal calf serum (FCS), 20 mM HEPES buffer and 100 μg/ml gentamicin. Curcumin was directly added to cell cultures at the indicated concentrations. Anti-ERK, anti-FADD, anti-Bcl-2, anti-PLC-γ1, anticaspase-3, anti-DR4, anti-DR5 and antiactin antibodies were purchased from Santa Cruz Biotechnology (Santa Cruz, CA). Anticaspase-8 and anticytochrome c were purchased form Stressgen (Stressgen, British Columbia, Canada). Soluble recombinant TRAIL was purchased from R&D Systems (Minneapolis, MN). Curcumin was obtained from Sigma Chemical Co.
Western blotting
Cellular lysates were prepared by suspending 1 × 10 6 cells in 100 μl of lysis buffer (137 mM NaCl, 15 mM EGTA, 0.1 mM sodium orthovanadate, 15 mM MgCl 2 , 0.1% Triton X-100, 25 mM MOPS, 100 μM phenylmethylsulfonyl fluoride and 20 μM leupeptin, adjusted to pH 7.2). The cells were disrupted by sonication and extracted at 4°C for 30 min. The proteins were electrotransferred to Immobilon-P membranes (Millipore, Bedford, MA). Detection of specific proteins was carried out with an ECL western blotting kit according to the manufacturer's instructions.
Cell count and flow cytometry analysis
Cell counts were performed using a hemocytometer. Approximately 1 × 10 6 Caki cells were suspended in 100 μl of phosphate buffered saline (PBS) and 200 μl of 95% ethanol was added while vortexing. The cells were incubated at 4°C for 1 h, washed with PBS, and resuspended in 250 μl of 1.12% sodium citrate buffer (pH 8.4) together with 12.5 μg of RNase. Incubation was continued at 37°C for 30 min. The cellular DNA was then stained by applying 250 μl of propidium iodide (50 μg/ml) for 30 min at room temperature. The stained cells were analyzed by fluorescent activated cell sorting (FACS) on a FACScan flow cytometer for relative DNA content based on red fluorescence.
DNA fragmentation assay
After treatment with curcumin, TRAIL, and combination of TRAIL and curcumin for 24 h, Caki cells were lysed in a buffer containing 10 mM Tris (pH 7.4), 150 mM NaCl, 5 mM EDTA and 0.5% Triton X-100 for 30 min on ice. Lysates were vortexed and cleared by centrifugation at 10000 g for 20 min. Fragmented DNA in the supernatant was extracted with an equal volume of neutral phenol:chloroform:isoamyl alcohol mixture (25:24:1) and analyzed electrophoretically on 2% agarose gels containing 0.1 µg/ml of ethidium bromide.
Asp-Glu-Val-Asp-ase (DEVDase) activity assay
To evaluate DEVDase activity, cell lysates were prepared after their respective treatment with TRAIL or curcumin. Assays were performed in 96-well microtiter plates by incubating 20 μg of cell lysates in 100 µl of reaction buffer [1% NP-40, 20 mM Tris–HCl (pH 7.5), 137 mM NaCl and 10% glycerol] containing the caspases substrate [Asp-Glu-Val-Asp-chromophore- p -nitroanilide (DVAD-pNA)] at 5 μM concentration. Lysates were incubated at 37°C for 2 h. Thereafter, the absorbance at 405 nm was measured with a spectrophotometer.
Measurement of reactive oxygen species
The intracellular accumulation of ROS was determined using the fluorescent probes DHE and H 2 DCFDA. DHE preferentially measures
Plasmids, transfections and luciferase gene assays
The human DR5 promoter (pDR5/SacI plasmid)-containing plasmids have been described previously ( 26 ). In brief, cells were plated onto 6-well plates at a density of 5 × 10 5 cells/well and grown overnight. Cells were cotransfected with 2 μg of various plasmid constructs and 1 μg of the pCMV-β-galactosidase plasmid for 5 h by the lipofectamine method. After transfection, cells were cultured in 10% fetal calf serum (FCS) medium with a vehicle (DMSO) or drugs for 24 h. Luciferase and β-galactosidase activities were assayed according to the manufacturer's protocol (Promega, Madison, WI). Luciferase activity was normalized for β-galactosidase activity in the cell lysate and expressed as an average of three independent experiments.
RNA isolation and reverse transcriptase–polymerase chain reaction
To determine whether the potential sensitizing effects of curcumin to TRAIL-mediated apoptosis were a result of increased levels of mRNA encoding DR5, we compared the levels of DR5 in Caki cells, which were treated with or without various concentrations of curcumin. DR5 mRNA expression was determined by reverse transcriptase–polymerase chain reaction (RT–PCR). Total cellular RNA was extracted from cells using the TRIzol reagent (Life Technologies). A cDNA was synthesized from 2 μg of total RNA using M-MLV reverse transcriptase (Gibco-BRL, Gaithersburg, MD). The cDNA for DR5 and actin were amplified by PCR using specific primers. The sequences of the sense and antisense primer for DR5 were 5′-AAGACC CTTGTGCTCGTTGT-3′ and 5′-GACACATTCGATGTCACTCCA-3′, respectively. PCR products were analyzed by agarose gel electrophoresis and visualized by using ethidium bromide.
Analysis of cytochrome c release
Cells (2 × 10 6 ) were harvested, washed once with ice-cold PBS and gently lysed for 2 min in 80 μl ice-cold lysis buffer [250 mM sucrose, 1 mM EDTA, 20 mM Tris–HCl (pH 7.2), 1 mM DTT, 10 mM KCl, 1.5 mM MgCl 2 , 5 μg/ml pepstatin A, 10 μg/ml leupeptin and 2 μg/ml aprotinin]. Lysates were centrifuged at 12000 g at 4°C for 10 min to obtain the supernatants (cytosolic extracts free of mitochondria) and the pellets (fraction that contains mitochondria). The resulting cytosolic fractions were used for western blot analysis with an anticytochrome c antibody.
Analysis of cell surface DR5
Cells were detached with 0.5 mM EDTA, and washed three times with PBS wash buffer supplemented with 0.5% BSA. Cells (5 × 10 5 ) were resuspended in 200 μl of PBS, stained with the primary antibody (1 μg/ml) and incubated for 30 min at 4°C. Unreacted antibody was removed by washing the cells twice with the same PBS buffer. Cells were stained with the secondary antibody conjugated with fluorescein isothiocyanate (FITC) and incubated for 30 min at 4°C. Unbound FITC-conjugated antibody was washed twice with PBS. Cells were resuspended in 200 μl of PBS. Cell-surface expression of DR5 receptor was determined by flow cytometry. Fluorescent intensity of the cells is directly proportional to the density of receptor.
Results
Curcumin sensitizes TRAIL-mediated apoptosis
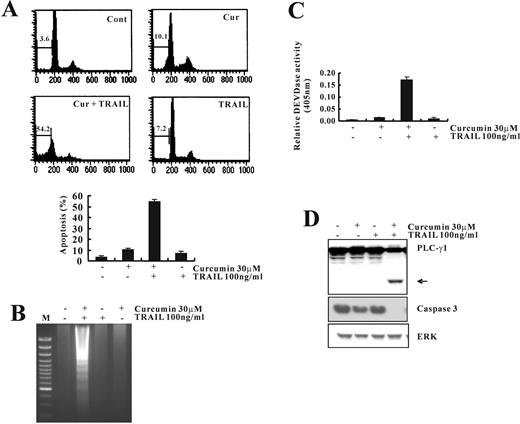
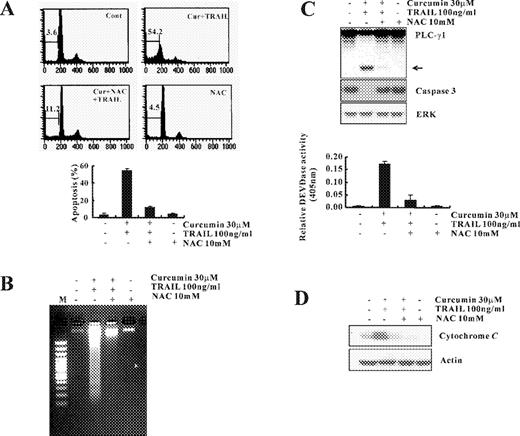
To investigate the effect of curcumin on TRAIL-mediated apoptosis, human renal cancer cell line Caki cells were treated with curcumin alone (30 μM), TRAIL alone (100 ng/ml), and with combination of curcumin and TRAIL. Three established criteria were subsequently used to assess apoptosis in our system. First, apoptosis in Caki cells was determined using flow cytometric analysis to detect hypodiploid cell populations. As shown in Figure 1A , cotreatment of Caki cells with curcumin and TRAIL resulted in a markedly increased accumulation of sub-G1 phase cells. Second, we analyzed DNA fragmentation, which is another hallmark of apoptosis. Following agarose gel electrophoresis of DNAs from Caki cells treated with curcumin and TRAIL for 24 h, a typical ladder pattern of internucleosomal fragmentation was observed. In contrast, DNA fragmentation in Caki cells treated with TRAIL alone or curcumin alone was barely detected ( Figure 1B ). In addition, we analyzed whether cotreatment with curcumin and TRAIL gave rise to the activation of caspases, a key executioner of apoptosis. Cotreatment of Caki cells with curcumin and TRAIL strongly stimulated DEVDase activity ( Figure 1C ) and led to a reduction of the protein levels of 32-kDa precursor together with a concomitant cleavage of phospholipase C-γ1 (PLC-γ1), a substrate protein of caspases ( Fig. 1D ). Taken together, these results indicate that treatment with curcumin sensitizes Caki cells to TRAIL-mediated apoptosis.
Curcumin sensitizes Caki cells to TRAIL-induced apoptosis. ( A ) Flow cytometric analysis of apoptotic cells. Caki cells were treated with TRAIL (100 ng/ml) in either the absence or the presence of curcumin (30 μM) for 24 h. Apoptosis was analyzed as a sub-G1 fraction by FACS. ( B ) Fragmentations of genomic DNA in Caki cells treated for 24 h with the indicated concentrations of curcumin and TRAIL. Fragmented DNA was extracted and analyzed on 2% agarose gel. ( C ) Cells were treated with the indicated concentrations of curcumin and TRAIL. Enzymatic activities of DEVDase were determined by incubation of 20 μg of total protein with 200 μM chromogenic substrate (DEVD-pNA) in a 100 μl assay buffer for 2 h at 37°C. The release of chromophore p -nitroanilide (pNA) was monitored spectrophotometrically (405 nm). ( D ) The expression levels of apoptosis-related proteins in Caki cells by treatment with curcumin and TRAIL. Equal amounts of cell lysates (40 μg) were resolved by SDS–PAGE, transferred to nitrocellulose membrane and probed with specific antibodies, anticaspase-3, anti-PLC-γ1 or with anti-ERK antibody to serve as control for the loading of protein level. The proteolytic cleavage of PLC-γ1 is indicated by an arrow. A representative study is shown; two additional experiments yielded similar results.
Curcumin upregulates DR5 in various cancer cells
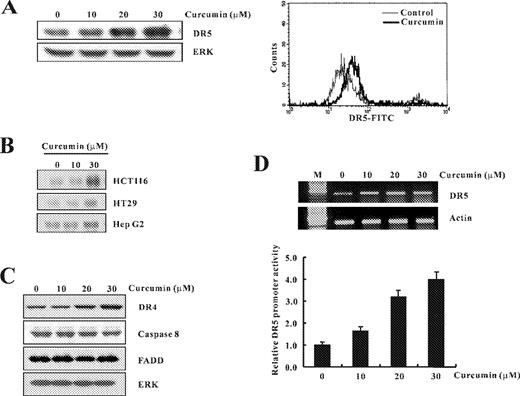
It is well known that TRAIL interacts with death receptors (DR4 and DR5) to trigger apoptotic signals. To assess the molecular mechanisms underlying the synergistic induction of apoptosis by curcumin and TRAIL in Caki cells, we first examined the effect of curcumin on the expression of DR5 by western blotting. As shown in Figure 2A , treatment with curcumin induced the expression of DR5 protein in a dose-dependent manner. Since transport of DR5 from the cytosol to the cell surface is crucial for TRAIL sensitivity ( 27 ), we confirmed the expression of DR5 receptor on the cell surface by flow cytometric analysis using specific DR5 antibodies. The expression levels of DR5 on the cell surface were markedly higher in cells treated with 20 μM curcumin than the untreated control ( Figure 2A ). Upregulation of DR5 proteins by curcumin was also observed in a variety of tumor cell types (colon cancer cells; HCT116 and HT29, and hepatocellular carcinoma HepG2 cells) ( Figure 2B ). We further analyzed the changes in various proteins related with TRAIL-mediated apoptotic signaling pathway ( 16 , 22 ) by Western blotting, following treatment of Caki cells with various concentrations of curcumin. As shown in Figure 2C , treatment with curcumin slightly induced the expression of DR4 protein in a dose-dependent manner. However, there were no major changes in the protein levels of caspase-8 or Fas associated death domain (FADD) ( Figure 2C ). Next, we investigated whether curcumin-mediated DR5 induction is controlled at the transcriptional level. RT–PCR analysis demonstrated that curcumin enhanced DR5 mRNA levels in a dose-dependent manner ( Figure 2D ). Moreover, transfection of Caki cells with the luciferase reporter construct containing DR5 promoter and the subsequent luciferase assay demonstrated that curcumin significantly increased DR5 promoter activity in a dose-dependent manner ( Figure 2D ).
The expression levels of the DR5 mRNA and DR5 protein by treatment with curcumin in Caki cells. ( A ) Caki cells were treated with the indicated concentrations of curcumin. Equal amounts of cell lysates (40 μg) were resolved by SDS–PAGE, transferred to nitrocellulose membrane and probed with specific antibodies, anti-DR5 or with anti-ERK antibody to serve as control for the loading of protein level. A representative study is shown; two additional experiments yielded similar results. Effects of curcumin on the cell-surface expression of DR5 in Caki cells. Caki cells were treated with 20 μM curcumin for 24 h. Expression levels of DR5 were measured by flow cytometric analysis. ( B ) HCT116, HT29 and HepG2 cells were treated with the indicated concentrations of curcumin. Expression of DR5 was determined by western blot analysis. ( C ) Effect of curcumin on TRAIL-mediated apoptosis related proteins. Caki cells were treated with the indicated concentrations of curcumin. Equal amounts of cell lysates (40 μg) were resolved by SDS–PAGE, transferred to nitrocellulose membrane and probed with specific antibodies, anti-DR4, anticaspase-8, anti-FADD or with anti-ERK antibody to serve as control for the loading of protein level. ( D ) Total RNA was isolated and RT–PCR analysis was performed as described in Materials and methods. A representative study is shown; two additional experiments yielded similar results. pDR5/SacI promoter plasmid was transfected, and treated with varying concentrations of curcumin. The cells were lysed and luciferase activity measured. Data represent the mean ± SD of at least three independent experiments.
Upregulation of DR5 by curcumin appears to be dependent on the formation of reactive oxygen metabolites
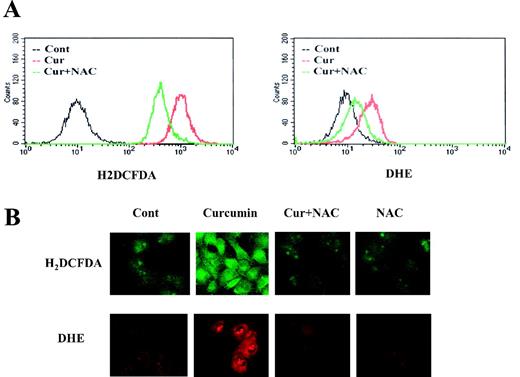
Many antineoplastic agents eliminate tumor cells by inducing programmed cell death or apoptosis, and numerous investigations have documented that oxidative stress-mediated cellular changes are frequently induced in cells exposed to cytotoxic drugs, UV or γ-irradiation ( 28 , 29 ). Therefore, we examined whether curcumin affects the cellular levels of peroxide and/or anion superoxides by measuring the changes in the fluorescence using H 2 DCFDA and DHE ( 24 , 25 ). As shown in Figure 3A , treatment with curcumin markedly increased the H 2 DCFDA- and DHE-derived fluorescence. This curcumin-mediated increase in fluorescence was markedly inhibited by pretreatment with NAC, an antioxidant. Production of ROS by curcumin was further confirmed using fluorescence microscopy. Caki cells treated with curcumin displayed intense fluorescence inside the cell on staining with H 2 DCFDA- or DHE-dye ( Figure 3B ), demonstrating the intracellular accumulation of ROS. This increased fluorescence derived from H 2 DCFDA or DHE in curcumin-treated Caki cells was also markedly reduced by NAC ( Figure 3B ).
Curcumin induces elevation of the intercellular ROS level. ( A ) To determine the intracellular content of peroxides and anion superoxide, Caki cells were loaded with H2DCFDA and DHE, respectively, and fluorescence was measured by flow cytometry. Caki cells were loaded with fluorescence-dye and further stimulated with curcumin (30 μM) in the presence or absence of NAC (5mM). After 4 h, H2DCFDA and DHE fluorescence was visualized using a confocal microscope ( B ).
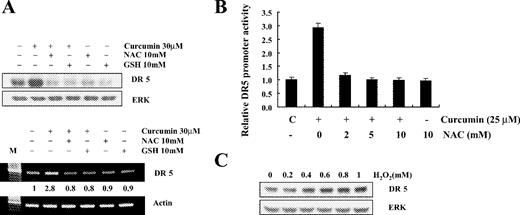
To investigate whether ROS generation is directly associated with curcumin-induced DR5 upregulation, we assessed the DR5 expression levels in cells treated with NAC or GSH in addition to 30 μM curcumin. As shown in Figure 4A , the treatment with curcumin significantly increased both the protein and mRNA levels of DR5, whereas pretreatment with NAC or GSH markedly inhibited this curcumin-induced DR5 upregulation. Curcumin-mediated increase in DR5 promoter activity, assessed by transient transfection and luciferase assay, was also blocked by pretreatment with NAC ( Figure 4B ). Involvement of ROS in DR5 upregulation was further examined by western blotting of DR5 in cells treated with H 2 O 2 . As shown in Figure 4C , treatment with H 2 O 2 induced DR5 protein levels in a dose-dependent manner. These data clearly indicate that ROS generation is critical for curcumin-induced DR5 upregulation.
Effects of NAC and GSH on curcumin-induced DR5 protein and mRNA expression. ( A ) Caki cells were incubated with indicated concentration of NAC and GSH for 1 h before challenge with curcumin (30 μM) for 24 h. Equal amounts of cell lysates (40 μg) were resolved by SDS–PAGE, transferred to nitrocellulose and probed with anti-DR5 antibodies. DR5 mRNA level was measured by RT–PCR analysis. The values below the figure represent change in mRNA expression of the bands normalized to actin. A representative study is shown; two additional experiments yielded similar results. ( B ) Caki cells were transfected with pDR5/SacI promoter plasmid and further cultured with curcumin (25 μM) in the presence or absence of NAC. The cells were lysed and luciferase activity measured. Data represent the mean ± SD of at least three independent experiments. ( C ) Caki cells were treated with the indicated concentrations of H 2 O 2 . DR5 protein expression level was determined by immunoblot analysis with anti-DR5 or with anti-ERK antibody to serve as control for the loading of protein level. A representative study is shown; two additional experiments yielded similar results.
Pretreatment with NAC, an antioxidant, prevents apoptosis by curcumin plus TRAIL
To examine the role of ROS in curcumin plus TRAIL-induced apoptosis, we used NAC, a thiol antioxidant, which is known to function as both a redox buffer and a reactive oxygen intermediate scavenger ( 30 ). Pretreatment of Caki cells with NAC markedly blocked curcumin plus TRAIL-induced apoptosis ( Figure 5A ). We also found that pretreatment with NAC prevented DNA fragmentation in cells treated with curcumin plus TRAIL ( Figure 5B ). These data clearly indicate that the blocking of curcumin plus TRAIL-induced apoptosis by NAC is associated with ROS generation. As shown in Figure 5C , NAC pretreatment significantly inhibited DEVDase activation and attenuated the cleavage of PLC-γ1 ( Figure 5C ). We next examined whether NAC has a direct effect on curcumin and TRAIL-mediated cytochrome c release, because the release of mitochondrial cytochrome c into the cytosol is upstream of caspases activation. Importantly, as shown in Figure 5D , pretreatment of Caki cells with NAC markedly blocked curcumin plus TRAIL-induced release of mitochondrial cytochrome c to the cytosol. These data clearly indicate that prevention of curcumin plus TRAIL-induced apoptosis by NAC is associated with the blocking of mitochondrial cytochrome c release and the subsequent activation of caspases.
Effect of NAC on curcumin plus TRAIL-induced apoptosis. ( A ) Caki cells were treated with curcumin plus TRAIL in the presence or absence of NAC (10 mM). Apoptosis was analyzed as a sub-G1 fraction by FACS. ( B ) Fragmentations of genomic DNA in Caki cells treated with curcumin plus TRAIL in the presence or absence of NAC (10 mM). Fragmented DNA was extracted and analyzed on 2% agarose gel. ( C ) Equal amounts of cell lysates (40 μg) were resolved by SDS–PAGE, transferred to nitrocellulose membrane and probed with specific antibodies, anticaspase-3, anti-PLC-γ1 or with anti-ERK antibody to serve as control for the loading of protein level. The proteolytic cleavage of PLC-γ1 is indicated by an arrow. A representative study is shown; two additional experiments yielded similar results. DEVDase activity was determined as described in Figure 1C . Data shown are means ± SD ( n = 3). ( D ) 30 μg of cytosolic protein was resolved on 12% SDS–PAGE and then transferred to nitrocellulose, and probed with specific anticytochrome c antibody or with antiactin antibody to serve as control for the loading of protein level.
Ectopic expression of Peroxiredoxin II suppresses curcumin-induced DR5 upregulation
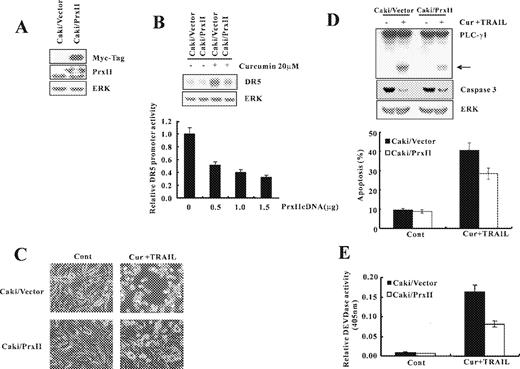
Prx II is an antioxidant enzyme that reduces the cellular levels of H 2 O 2 and other reactive oxygen species using thioredoxin as the immediate electron donor and its peroxidase activity prevents cells from ROS insult ( 31 , 32 ). To further evaluate the effect of ROS signaling on curcumin-mediated DR5 regulation and apoptosis induced by curcumin plus TRAIL, we established control cells (Caki/vector cells) and Caki/Prx II cells by stable transfection of Caki with control vector and the plasmid encoding Myc-tagged form of Prx II, respectively ( Figure 6A ). Expression of Prx II significantly reduced DR5 protein levels and DR5 promoter activity after treatment with curcumin ( Figure 6B ). While treatment with curcumin plus TRAIL, induced significant apoptosis in Caki/vector cells, the apoptotic characteristics, such as cell shrinkage, apoptotic bodies and detachment from the plates, were much less frequently observed in Caki/Prx II cells treated with curcumin plus TRAIL ( Figure 6C ). Moreover, ectopic expression of Prx II significantly inhibited reduction of the protein levels of 32-kDa precursor and cleavage of PLC-γ1, a substrate protein of caspases ( Figure 6D ). In parallel with the degradation of caspase-3, after 24 h exposure to curcumin plus TRAIL, Caki/vector cells exhibited a significant increase in DEVDase activity, whereas only a half level of the activity was observed in Caki/Prx II cells ( Figure 6E ). Taken together, Prx II overexpression protects Caki cells against curcumin plus TRAIL-induced apoptosis.
Ectopic expression of Prx II reduced curcumin plus TRAIL-induced apoptosis. ( A ) Whole cell lysates obtained from Caki cells stably transfected with a Prx II-expression vector or the empty vector were subjected to SDS–PAGE, transferred to membranes, and immunoblotted using anti-Prx II and anti-Myc antibodies as indicated. ( B ) Caki/Prx II and Caki/Vector cells were treated with curcumin (20 μM). DR5 protein expression level was determined by immunoblot analysis with anti-DR5 or with anti-ERK antibody to serve as control for the loading of protein level. A representative study is shown; two additional experiments yielded similar results. pDR5/SacI promoter plasmid was co-transfected with varying concentrations of Prx II plasmid. The cells were lysed and luciferase activity measured. Data represent the mean ± SD of at least three independent experiments. ( C ) Caki/Prx II and Caki/Vector cells were treated with a vehicle and curcumin plus TRAIL for 24 h. Magnification, 200×. ( D ) Equal amounts of cell lysates (40 μg) were resolved by SDS–PAGE, transferred to nitrocellulose membrane, and probed with anticaspase-3 and anti-PLC-γ1 antibodies. The proteolytic cleavage of PLC-γ1 is indicated by an arrow. A representative study is shown; two additional experiments yielded similar results. ( E ) DEVDase activity was determined as described in Figure 1C . Data shown are means ± SD ( n = 3).
Apoptosis induced by curcumin plus TRAIL is not blocked by Bcl-2-overexpression
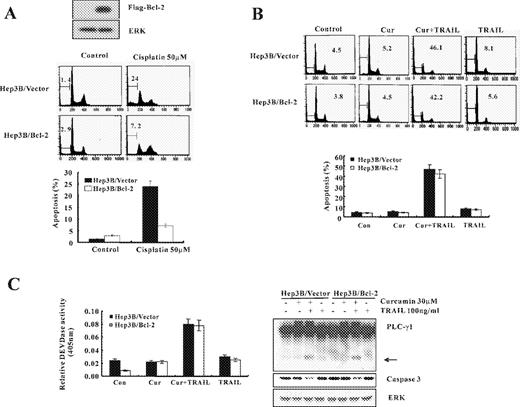
Overexpression of Bcl-2 is known to inhibit chemotherapy-induced apoptosis ( 33 ). To determine the effect of cotreatment with curcumin and TRAIL on the viability of Bcl-2 overexpressing cells, we employed Hep3B hepatocellular carcinoma cells engineered for overexpression of Bcl-2 (Hep3B/Bcl-2) and vector-transfected control cells (Hep3B/Vector) ( Figure 7A ). To confirm antiapoptotic role of Bcl-2, vector and Bcl-2 overexpressing cells were treated with an anticancer drug, cisplatin. As shown in Figure 7A , treatment of Hep3B/vector cells with 50 μM cisplatin resulted in a markedly increased accumulation of sub-G1 phase, but not in Hep3B/Bcl-2 cells. Cells were treated for 24 h with curcumin alone (30 μM), TRAIL alone (100 ng/ml), and combination of curcumin and TRAIL. As shown in Figure 7B , similar percentages of the apoptotic cells were detected in control cells and Bcl-2 overexpressing cells treated with curcumin plus TRAIL.
Curcumin plus TRAIL induces apoptotic cell death in Bcl-2 overexpressing cells. ( A ) Antiapoptotic effect of Bcl-2 on cisplatin-induced apoptosis. Analysis of Bcl-2 expression in the stably transfected cell lines. Western blotting using an anti-Flag antibody was performed to detect Flag-tagged Bcl-2 in the selected cell lines. Hep 3B/Vector and Hep 3B/Bcl-2 cells were treated with or without 50 μM cisplatin, and apoptosis was analyzed as a sub-G1 fraction by FACS. Data are mean values obtained from three independent experiments and bars represent standard deviations. ( B ) Hep 3B/Vector and Hep 3B/Bcl-2 cells were treated for 24 h with curcumin alone (30 μM), TRAIL alone (100 ng/ml), and combination of curcumin and TRAIL. Apoptosis was analyzed as a sub-G1 fraction by FACS. ( C ) Equal amounts of cell lysates (40 μg) were subjected to electrophoresis and analyzed by western blot for caspase-3 and PLC-γ1. The proteolytic cleavage of PLC-γ1 is indicated by an arrow. DEVDase activity was determined as described in Figure 1C . Data shown are means ± SD ( n = 3).
In addition, ectopic expression of Bcl-2 did not inhibit reduction of the protein levels of 32-kDa precursor and cleavage of PLC-γ1 ( Figure 7C ). After 24 h exposure to curcumin plus TRAIL, both Caki/vector and Caki/Bcl-2 cells exhibited a similar increase in DEVDase activity ( Figure 7C ). These data suggest that the combined treatment with curcumin and TRAIL may effectively induce apoptotic cell death in Bcl-2 overexpressing cancer cells, which are relatively resistant to chemotherapy.
Discussion
TRAIL is a recently identified member of the TNF family and is capable of inducing apoptosis in various tumor cells ( 16 , 19 ). TRAIL induces apoptosis by interacting with two death-inducing receptors, DR4 and DR5. The death receptors DR4 and/or DR5 have been demonstrated to play a crucial role in synergistic cytotoxicity associated with TRAIL and chemotherapeutic agents ( 16 , 19 ). Several reports have also shown that chemotherapeutic agents and ionizing radiation can sensitize TRAIL-induced cytotoxicity by decreasing intracellular levels of Flice-inhibitory protein (FLIP) ( 34 , 35 ). FLIP structurally resembles caspase-8 but lacks proteolytic activity, thus functioning as a dominant negative inhibitor of caspase-8 ( 36 – 38 ).
We demonstrate in the present study that subtoxic concentrations of the curcumin sensitize human renal cancer cells to TRAIL-mediated apoptosis, although we have previously shown that high concentrations of curcumin induce apoptosis in these cells ( 10 ). Recently, Deeb et al . ( 39 ) reported that curcumin sensitizes prostate cancer cells to TRAIL by inhibiting NF-κB through suppression of IκBα phosphorylation ( 40 ). However, in this study we provide a novel mechanism that shows the upregulation of DR5 is essential for curcumin plus TRAIL-induced apoptosis.
Recently it has been suggested that oxidative stress plays a role as a common mediator of apoptosis ( 41 ). We and other groups reported that curcumin generates reactive oxygen intermediates in cancer cells ( 9 , 10 ). Accumulation of intercellular ROS leads to disruption of the mitochondrial membrane potential, release of cyctochrome c into the cytosol with subsequent activation of the caspase cascade, and ultimately to programmed cell death through apoptosis ( 10 ). In this study, we demonstrate that not only curcumin-induced ROS but also H 2 O 2 can induce DR5 upregulation through transcriptional control, although it remains to be clarified how ROS increase the transcriptional activity of DR5 promoter. The antioxidant NAC prevents both the curcumin-mediated DR5 upregulation and apoptosis induced by curcumin plus TRAIL. Prx II protein has been proposed to play a role as a thiol-specific antioxidative protein ( 31 , 32 ) that reacts with thiol-containing proteins such as thioredoxin. Prx II, which is induced by various oxidative stimuli, has been shown to play an important protective role in oxidative damage by H 2 O 2 ( 42 ). In our study, ectopic expression of Prx II also reduced curcumin plus TRAIL-induced apoptosis, downregulating DR5 expression. Taken together, these results demonstrate a critical role of ROS-mediated DR5 upregulation in curcumin-stimulated TRAIL-mediated apoptosis. Our results provide the first mechanistic evidence that curcumin treatment results in upregulation of DR5 and renders cells more sensitive to the cytotoxic activities of TRAIL.
Clinically, resistance to apoptosis is a major obstacle in chemotherapeutic treatments of cancers. The apoptosis inducing ability of curcumin, in conjunction with its non-toxic nature, could make it a potentially effective preventive and/or therapeutic agent against tumor. Moreover, the combination of curcumin and TRAIL demonstrated considerable cytotoxicity in tumor overexpressing Bcl-2, which confers general resistance to many other chemotherapeutic treatments. Thus, in terms of a clinical perspective, the combination of curcumin and TRAIL may be a novel strategy for the treatment of a variety of human cancers that are resistant to chemotherapy. However, the potential clinical implications of our studies will depend on whether or not curcumin can be given safely to humans at doses high enough to achieve pharmacologically active levels. Clinical trials of oral curcumin cannot be utilized because of poor bioavailability due to its rapid metabolism in liver and intestinal wall. However, coingestion of curcumin with the pepper constituent l-piperoylpiperidine, which is used to inhibit xenobiotic glucuronidation, appeared to increase curcumin serum AUC by a factor of 20 ( 43 ). In addition, a recent clinical pilot study in colorectal cancer suggests that curcuma extract can be administered safely to patients at doses of up to 2.2 g daily, equivalent to 180 mg of curcumin ( 44 ). Therefore, additional in vivo studies are needed to evaluate the applicability of curcumin as a chemopreventive and/or therapeutic agent for cancer.
Both authors contributed equally to this work.
This work was supported by the Korea Science and Engineering Foundation (KOSEF) through the MRC at Keimyung University (R13-2002-028-03001-0) and by grant No. R01-2005-000-10786-0 from MOST (KOSEF).
Conflict of Interest Statement : None declared.
References
Mehta,K., Pantazis,P., McQueen,T. and Aggarwal,B.B. (
Huang,M.T., Smart,R.C., Wong,C.Q. and Conney,A.H. (
Huang,M.T., Ma,W., Lu,Y.P., Chang,R.L., Fisher,C., Manchand,P.S., Newmark,H.L. and Conney,A.H. (
Huang,M.T., Ma,W., Yen,P., Xie,J.G., Han,J., Frenkel,K., Grunberger,D. and Conney,A.H. (
Huang,M.T., Lou,Y.R., Ma,W., Newmark,H.L., Reuhl,K.R. and Conney,A.H. (
Kelloff,G.J., Boone,C.W., Crowell,J.A., Steele,V.E., Lubet,R. and Sigman,C.C. (
Somasundaram,S., Edmund,N.A., Moore,D.T., Small,G.W., Shi,Y.Y. and Orlowski,R.Z. (
Jaiswal,A.S., Marlow,B.P., Gupta,N. and Narayan,S. (
Bhaumik,S., Anjum,R., Rangaraj,N., Pardhasaradhi,B.V. and Khar,A. (
Woo,J.H., Kim,Y.H., Choi,Y.J. et al . (
Kehrer,J.P. (
Shiraishi,F., Curtis,L.M., Truong,L., Poss,K., Visner,G.A., Madsen,K., Nick,H.S. and Agarwal,A. (
Zhang,X.D., Nguyen,T., Thomas,W.D., Sanders,J.E. and Hersey,P. (
Dejosez,M., Ramp,U., Mahotka,C., Krieg,A., Walczak,H., Gabbert,H.E. and Gerharz,C.D. (
Srivastava,R.K. (
Pan,G., O'Rourke,K., Chinnaiyan,A.M., Gentz,R., Ebner,R., Ni,J. and Dixit,V.M. (
Pan,G., Ni,J., Wei,Y.F., Yu,G., Gentz,R. and Dixit,V.M. (
Ashkenazi,A. and Dixit,V.M. (
Sheridan,J.P., Marsters,S.A., Pitti,R.M. et al . (
Ozoren,N., Fisher,M.J., Kim,K., Liu,C.X., Genin,A., Shifman,Y., Dicker,D.T., Spinner,N.B., Lisitsyn,N.A. and El-Deiry,W.S. (
Wang,S. and El-Deiry,W.S. (
Ivanov,V.N., Bhoumik,A. and Ronai,Z. (
Benov,L., Sztejnberg,L. and Fridovich,I. (
LeBel,C.P., Ischiropoulos,H. and Bondy,S.C. (
Kim,Y.H., Park,J.W., Lee,J.Y. and Kwon,T.K. (
Jin,Z., McDonald,E.R., Dicker,D.T. and El-Deiry,W.S. (
Thompson,C.B. (
Jacobson,M.D. (
Rhee,S.G., Kang,S.W., Netto,L.E., Seo,M.S. and Stadtman,E.R. (
Wood,Z.A., Schroder,E., Robin,H.J. and Poole,L.B. (
Johnstone,R.W., Ruefli,A.A. and Lowe,S.W. (
Martelli,A.M., Tazzari,P.L., Tabellini,G., Bortul,R., Billi,A.M., Manzoli,L., Ruggeri,A., Conte,R. and Cocco,L. (
Shankar,S. and Srivastava,R.K. (
Thome,M., Schneider,P., Hofmann,K. et al . (
Griffith,T.S., Chin,W.A., Jackson,G.C., Lynch,D.H. and Kubin,M.Z. (
Scaffidi,C., Schmitz,I., Krammer,P.H. and Peter,M.E. (
Deeb,D., Xu,Y.X., Jiang,H., Gao,X., Janakiraman,N., Chapman,R.A. and Gautam,S.C. (
Deeb,D., Jiang,H., Gao,X., Hafner,M.S., Wong,H., Divine,G., Chapman,R.A., Dulchavsky,S.A. and Gautam,S.C. (
Simon,H.U., Haj-Yehia.A. and Levi-Schaffer,F. (
Kim,H., Lee,T.H., Park,E.S., Suh,J.M., Park,S.J., Chung,H.K., Kwon,O.Y., Kim,Y.K., Ro,H.K. and Shong,M. (
Shoba,G., Joy,D., Joseph,T., Majeed,M., Rajendran,R. and Srinivas,P.S. (
Author notes
Department of Immunology, School of Medicine, Keimyung University, 194 DongSan-Dong Jung-Gu, Taegu 700-712, South Korea and 1Institute for Medical Sciences, Ajou University School of Medicine, 5 Woncheon-Dong, Paldal-Gu, Suwon 442-749, South Korea