Abstract
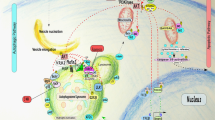
Autophagy regulates cell death both positively and negatively, but the molecular basis for this paradox remains inadequately characterized. We demonstrate here that transient cell-to-cell variations in autophagy can promote either cell death or survival depending on the stimulus and cell type. By separating cells with high and low basal autophagy using flow cytometry, we demonstrate that autophagy determines which cells live or die in response to death receptor activation. We have determined that selective autophagic degradation of the phosphatase Fap-1 promotes Fas apoptosis in Type I cells, which do not require mitochondrial permeabilization for efficient apoptosis. Conversely, autophagy inhibits apoptosis in Type II cells (which require mitochondrial involvement) or on treatment with TRAIL in either Type I or II cells. These data illustrate that differences in autophagy in a cell population determine cell fate in a stimulus- and cell-type-specific manner. This example of selective autophagy of an apoptosis regulator may represent a general mechanism for context-specific regulation of cell fate by autophagy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Mizushima, N., Levine, B., Cuervo, A.M. & Klionsky, D.J. Autophagy fights disease through cellular self-digestion. Nature 451, 1069–1075 (2008).
Kroemer, G., Marino, G. & Levine, B. Autophagy and the integrated stress response. Mol. Cell 40, 280–293 (2010).
Maycotte, P. & Thorburn, A. Autophagy and cancer therapy. Cancer Biol. Ther. 11, 127–137 (2011).
White, E. J. et al. Autophagy regulation in cancer development and therapy. Am. J. Cancer Res. 1, 362–372 (2011).
Dikic, I., Johansen, T. & Kirkin, V. Selective autophagy in cancer development and therapy. Cancer Res. 70, 3431–3434 (2010).
White, E. Deconvoluting the context-dependent role for autophagy in cancer. Nat. Rev. Cancer 12, 401–410 (2012).
Choi, A. M., Ryter, S. W. & Levine, B. Autophagy in human health and disease. New Engl. J. Med. 368, 651–662 (2013).
Gump, J. M. & Thorburn, A. Autophagy and apoptosis: what is the connection? Trends Cell Biol. 21, 387–392 (2011).
Eisenberg-Lerner, A., Bialik, S., Simon, H. U. & Kimchi, A. Life and death partners: apoptosis, autophagy and the cross-talk between them. Cell Death Differ. 16, 966–975 (2009).
Maiuri, M.C., Zalckvar, E., Kimchi, A. & Kroemer, G. Self-eating and self-killing: crosstalk between autophagy and apoptosis. Nat. Rev. Mol. Cell Biol. 8, 741–752 (2007).
Yonekawa, T. & Thorburn, A. Autophagy and cell death. Essays Biochem. 55, 105–117 (2013).
Norman, J. M., Cohen, G. M. & Bampton, E. T. The in vitro cleavage of the hAtg proteins by cell death proteases. Autophagy 6, 1042–1056 (2010).
Luo, S. et al. Bim inhibits autophagy by recruiting Beclin 1 to microtubules. Mol. Cell 47, 359–370 (2012).
Wirawan, E. et al. Caspase-mediated cleavage of Beclin-1 inactivates Beclin-1-induced autophagy and enhances apoptosis by promoting the release of proapoptotic factors from mitochondria. Cell Death Dis. 1, e18 (2010).
Radoshevich, L. et al. ATG12 conjugation to ATG3 regulates mitochondrial homeostasis and cell death. Cell 142, 590–600 (2010).
Rubinstein, A. D., Eisenstein, M., Ber, Y., Bialik, S. & Kimchi, A. The autophagy protein Atg12 associates with antiapoptotic Bcl-2 family members to promote mitochondrial apoptosis. Mol. Cell 44, 698–709 (2011).
Yousefi, S. et al. Calpain-mediated cleavage of Atg5 switches autophagy to apoptosis. Nat. Cell Biol. 8, 1124–1132 (2006).
Jin, Z. et al. Cullin3-based polyubiquitination and p62-dependent aggregation of caspase-8 mediate extrinsic apoptosis signaling. Cell 137, 721–735 (2009).
Hou, W., Han, J., Lu, C., Goldstein, L. A. & Rabinowich, H. Autophagic degradation of active caspase-8: a crosstalk mechanism between autophagy and apoptosis. Autophagy 6, 891–900 (2010).
Young, M. M. et al. Autophagosomal membrane serves as platform for intracellular death-inducing signaling complex (iDISC)-mediated caspase-8 activation and apoptosis. J. Biol. Chem. 287, 12455–12468 (2012).
Fullgrabe, J. et al. The histone H4 lysine 16 acetyltransferase hMOF regulates the outcome of autophagy. Nature 500, 468–471 (2013).
McPhee, C. K., Logan, M. A., Freeman, M. R. & Baehrecke, E. H. Activation of autophagy during cell death requires the engulfment receptor Draper. Nature 465, 1093–1096 (2010).
Yu, L. et al. Autophagic programmed cell death by selective catalase degradation. Proc. Natl Acad. Sci. USA 103, 4952–4957 (2006).
Gupta, P. B. et al. Stochastic state transitions give rise to phenotypic equilibrium in populations of cancer cells. Cell 146, 633–644 (2011).
Kreso, A. et al. Variable clonal repopulation dynamics influence chemotherapy response in colorectal cancer. Science 339, 543–548 (2013).
Gaudet, S., Spencer, S. L., Chen, W. W. & Sorger, P. K. Exploring the contextual sensitivity of factors that determine cell-to-cell variability in receptor-mediated apoptosis. PLoS Comput. Biol. 8, e1002482 (2012).
Guo, J. Y. et al. Activated Ras requires autophagy to maintain oxidative metabolism and tumorigenesis. Genes Dev. 25, 460–470 (2011).
Kaminskyy, V. O., Piskunova, T., Zborovskaya, I. B., Tchevkina, E. M. & Zhivotovsky, B. Suppression of basal autophagy reduces lung cancer cell proliferation and enhances caspase-dependent and -independent apoptosis by stimulating ROS formation. Autophagy 8, 1032–1044 (2012).
Hundeshagen, P., Hamacher-Brady, A., Eils, R. & Brady, N. R. Concurrent detection of autolysosome formation and lysosomal degradation by flow cytometry in a high-content screen for inducers of autophagy. BMC Biol. 9, 38 (2011).
Kimura, S., Noda, T. & Yoshimori, T. Dissection of the autophagosome maturation process by a novel reporter protein, tandem fluorescent-tagged LC3. Autophagy 3, 452–460 (2007).
Eng, K. E., Panas, M. D., Hedestam, G. B. & McInerney, G. M. A novel quantitative flow cytometry-based assay for autophagy. Autophagy 6, 634–641 (2010).
Sato, T., Irie, S., Kitada, S. & Reed, J. C. FAP-1: a protein tyrosine phosphatase that associates with Fas. Science 268, 411–415 (1995).
Li, Y. et al. Negative regulation of Fas-mediated apoptosis by FAP-1 in human cancer cells. Int. J. Cancer 87, 473–479 (2000).
Ivanov, V. N. et al. FAP-1 association with Fas (Apo-1) inhibits Fas expression on the cell surface. Mol. Cell Biol. 23, 3623–3635 (2003).
Ivanov, V. N., Ronai, Z. & Hei, T. K. Opposite roles of FAP-1 and dynamin in the regulation of Fas (CD95) translocation to the cell surface and susceptibility to Fas ligand-mediated apoptosis. J. Biol. Chem. 281, 1840–1852 (2006).
Takamura, A. et al. Autophagy-deficient mice develop multiple liver tumors. Genes Dev. 25, 795–800 (2011).
Ying, J. et al. Epigenetic disruption of two proapoptotic genes MAPK10/JNK3 and PTPN13/FAP-1 in multiple lymphomas and carcinomas through hypermethylation of a common bidirectional promoter. Leukemia 20, 1173–1175 (2006).
Yeh, S. H. et al. Genetic characterization of fas-associated phosphatase-1 as a putative tumor suppressor gene on chromosome 4q21.3 in hepatocellular carcinoma. Clin. Cancer Res. 12, 1097–1108 (2006).
Schickel, R., Park, S. M., Murmann, A. E. & Peter, M. E. miR-200c regulates induction of apoptosis through CD95 by targeting FAP-1. Mol. Cell 38, 908–915 (2010).
Barnhart, B. C., Alappat, E. C. & Peter, M. E. The CD95 type I/type II model. Semin. Immunol. 15, 185–193 (2003).
Algeciras-Schimnich, A. et al. Molecular ordering of the initial signaling events of CD95. Mol. Cell Biol. 22, 207–220 (2002).
Schmitz, I., Walczak, H., Krammer, P. H. & Peter, M. E. Differences between CD95 type I and II cells detected with the CD95 ligand. Cell Death Differ. 6, 821–822 (1999).
Meinhold-Heerlein, I. et al. Expression and potential role of Fas-associated phosphatase-1 in ovarian cancer. Am. J. Pathol. 158, 1335–1344 (2001).
Moscat, J. & Diaz-Meco, M. T. p62 at the crossroads of autophagy, apoptosis, and cancer. Cell 137, 1001–1004 (2009).
Kraft, C., Peter, M. & Hofmann, K. Selective autophagy: ubiquitin-mediated recognition and beyond. Nat. Cell Biol. 12, 836–841 (2010).
Paul, S., Kashyap, A. K., Jia, W., He, Y. W. & Schaefer, B. C. Selective autophagy of the adaptor protein Bcl10 modulates T cell receptor activation of NF-kappaB. Immunity 36, 947–958 (2012).
Ichimura, Y. & Komatsu, M. Selective degradation of p62 by autophagy. Semin. Immunopathol. 32, 431–436 (2010).
Kirkin, V., McEwan, D. G., Novak, I. & Dikic, I. A role for ubiquitin in selective autophagy. Mol. Cell 34, 259–269 (2009).
Levine, B. & Yuan, J. Autophagy in cell death: an innocent convict? J. Clin. Invest. 115, 2679–2688 (2005).
Zhu, J. H. et al. Protein tyrosine phosphatase PTPN13 negatively regulates Her2/ErbB2 malignant signaling. Oncogene 27, 2525–2531 (2008).
Amaravadi, R. K. et al. Principles and current strategies for targeting autophagy for cancer treatment. Clin. Cancer Res. 17, 654–666 (2011).
Klionsky, D. J. et al. Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy 8, 445–544 (2012).
Acknowledgements
We thank all members of the A.T. laboratory for thoughtful discussions and critical comments. We are grateful to K. Helm, L. Acosta and C. Childs of the University of Colorado Cancer Center Flow Cytometry Core for their guidance and assistance. We also thank D. Dill and the University of Colorado Anschutz Medical Campus (UCAMC) Electron Microscopy Core for electron micrographs and technical assistance, and R. Moldovan at the UCAMC Advanced Light Microscopy Core for assistance with confocal imaging. This work was supported by National Institutes of Health grants R01 CA111421 and CA150925 (A.T.), HL68628 (D.W.H.R.), and Shared Resources supported by P30 CA46934. J.M.G. was previously supported by 5T32CA82086-10 (UCAMC Department of Pediatrics) and is now an American Cancer Society Postdoctoral Fellow.
Author information
Authors and Affiliations
Contributions
J.M.G. and A.T. designed the study; J.M.G. and L.S. performed experiments; A.B. and D.W.H.R. provided Fap-1 reagents and expertise; J.M.G. designed experiments and analysed data; J.M.G. and A.T. wrote the manuscript, which was commented on by all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Method to sort cells with high and low autophagic flux by flow cytometry accurately measures basal and induced autophagic flux.
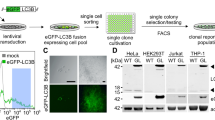
a, Cells constitutively expressing mCherry-GFP-LC3 are sorted based on the relative ratio of mCherry/EGFP fluorescence which changes in response to the pH gradient as autophagosomes fuse with lysosomes to form autolysosomes (autophagic flux). b, BJAB mCherry-GFP-LC3 cells were treated with EBSS (100%) or trehalose (75 mM) for 4 hours followed by flow cytometry for autophagic flux. Data are representative of at least 3 independent experiments. c, HeLa mCherry-GFP-LC3 cells expressing control or Atg5 shRNA were treated with EBSS for 4 hours followed by flow cytometry for autophagic flux. Data are representative of at least 3 independent experiments. d, BJAB mCherry-GFP-LC3 cells were sorted by flow cytometry for high or low autophagic flux, then treated with Lysotracker Blue for 30 min. at 37 °C, followed by flow cytometry (median GFP, mCherry or Lysotracker Blue fluorescence normalized to fold over unsorted control, mean ± s.e.m., n = 3 wells). e, BJAB mCherry-GFP-LC3 cells were sorted as in (d) followed by cytoplasmic extraction with 0.1 % saponin to eliminate non-lipidated LC3 (ref. 1) and flow cytometry to quantitate autophagosome/autolysosome number (normalized fold over unsorted control median GFP or mCherry fluorescence, mean ± s.e.m., n = 3 wells). f, Flow cytometry histograms of representative data from (e).
Supplementary Figure 2 Confocal microscopy reveals differences in the number of autophagosomes and autolysosomes in flux sorted cells; High and low flux sorted cells exhibit similar size, cell cycle and apoptotic profile.
a, HeLa mCherry-GFP-LC3 cells were sorted for autophagic flux, treated with Lysotracker Blue for 30 min. at 37 °C, adhered to slides by cytospin centrifugation, fixed with formaldehyde and visualized by spinning disc confocal microscopy; these are the same fields depicted in Fig. 1d. Colocalization images were produced using ImageJ (see methods). b, Quantification of the colocalization images in (a). Data are representative of 20 separate fields and have been repeated 5 times. c, BJAB mCherry-GFP-LC3 cells were sorted for autophagic flux by flow cytometry. d, e, Forward and side scatter of sorted cells in (c). f, Cell cycle profiles of sorted cells in (c) stained with Hoechst 33342.
Supplementary Figure 3 Autophagy modulation of Fas ligand-induced apoptosis and cell killing.
a, b, BJAB mCherry-GFP-LC3 cells were sorted for autophagic flux by flow cytometry followed by treatment with Fas ligand (a) or TRAIL (b) for 24 hours and MTS assay for viability (fold over no ligand control, mean ± s.e.m., n = 3 wells). These are the full data sets from the same experiment depicted in Figure 2d. c-e, BJAB mCherry-GFP-LC3 cells were sorted as in (a) followed by treatment with Fas ligand for the indicated times in the presence of fluorogenic Caspase-3/7 substrate CellEvent Green (5 μM) and analyzed by flow cytometry. c, Dot plot of Caspase-3/7 activity vs. DAPI fluorescence in unsorted cells. d, Quantitation of Caspase-3/7 activity in sorted cells at the indicated time points (median CellEvent Green fluorescence, mean ± s.e.m., n = 3 wells, *p = 0.0091). e, Representative flow cytometry histograms of Caspase-3/7 activity in (c, d). f, BJAB mCherry-GFP-LC3 cells were sorted for autophagic flux as above, followed by treatment with Fas ligand at the indicated concentrations in the presence of fluorogenic CellEvent Caspase-3/7 substrate (5 μM) and analyzed on an IncuCyte ZOOM for the indicated times (mean CellEvent Green fluorescence per field, mean, n = 12: 4 fields for each of 3 wells).
Supplementary Figure 4 Effect of autophagy inhibition or autophagy induction on Fas ligand-induced death.
a, BJAB cells expressing control or Atg12 shRNA were treated with Fas ligand (15ng/mL) for 24 hours in the presence or absence of zVAD-fmk (100 μM). Cell viability was then quantitated by MTS absorbance (fold over untreated control, mean ± s.e.m., n = 3 wells). b, BJAB cells expressing control or Bid shRNA were treated with autophagy inhibitor chloroquine (25 μM) for 12 hours followed by Fas ligand (4ng/mL) for 24 hours. Cell viability was then quantitated by MTS (fold over untreated control, mean ± s.e.m., n = 3 wells). c, BJAB cells were treated with autophagy inhibitors chloroquine (20 μM)or desmethylclomipramine (DCMI, 10 μM) for 12 hours followed by treatment with the indicated cytotoxic agents for 24 hours. Cells were then assayed for viability by MTS (% of control (no cytotoxic drug), mean ± s.e.m., n = 3 wells, *p = 0.0024,**p = 0.017). d, BJAB cells expressing vector control or the indicated autophagy dominant-negative constructs were treated with Fas ligand (4 ng/mL) for 24 hours; cell viability was then quantitated by MTS absorbance (fold over untreated control, mean ± s.e.m., n = 3 wells). Data are representative of 2 independent experiments. e, BJAB cells expressing control or Atg5 shRNA were treated with chloroquine (20 μM) for 12 hours followed by Fas ligand (4 ng/mL) for 24 hours. Cell viability was then quantitated by MTS (fold over no ligand control, mean ± s.e.m., n = 3 wells). Data are representative of 3 independent experiments. f, g, BJAB cells were treated with the indicated autophagy inducers overnight (EBSS, 100%; trehalose, 75 mM) followed by treatment with Fas ligand at the indicated concentrations for 24 hours. Cell viability was then quantitated by MTS absorbance. f, MTS data normalized (fold over untreated (no Fas ligand) control, mean ± s.e.m., n = 3 wells). g, Raw MTS values (absorbance at 490 nm, mean ± s.e.m., n = 3 wells).
Supplementary Figure 5 Autophagy controls cell surface expression of Fas receptor via Fap-1.
a, BJAB mCherry-GFP-LC3 cells were sorted for autophagic flux by flow cytometry, and then stained with APC-conjugated anti-Fas antibody at 4 °C for 30 minutes. Cells were then washed and analyzed by flow cytometry. Data are representative of 3 replicates from 3 independent experiments. b, Additional exposures of immunoblots from experiment in Fig. 4f. c-d, BJAB cells expressing control or Fap-1 shRNAs were treated with 20 μM chloroquine for 12 hours followed by treatment with Fas ligand (c) or TRAIL (d) at the indicated concentrations for 24 hours. Cell viability was quantitated by MTS (absorbance at 490 nm, mean ± s.e.m., n = 3 wells). These are the full dose response curves (without normalization) from the data in Figure 5c. e, Additional exposures from immunoprecipitation experiment depicted in Fig. 6c.
Supplementary Figure 6 Full immunoblot images.
Red boxes indicate the cropped portion of each western blot presented in the corresponding main figures.
Supplementary information
Supplementary Information
Supplementary Information (PDF 2390 kb)
Rights and permissions
About this article
Cite this article
Gump, J., Staskiewicz, L., Morgan, M. et al. Autophagy variation within a cell population determines cell fate through selective degradation of Fap-1. Nat Cell Biol 16, 47–54 (2014). https://doi.org/10.1038/ncb2886
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2886
This article is cited by
Autophagy: Regulator of cell death
Cell Death & Disease (2023)
Tight association of autophagy and cell cycle in leukemia cells
Cellular & Molecular Biology Letters (2022)
DNAJC24 is a potential therapeutic target in hepatocellular carcinoma through affecting ammonia metabolism
Cell Death & Disease (2022)
Lymphocytes are less sensitive to autophagy than monocytes during fasting and exercise conditions
Apoptosis (2022)
Autophagy blockade synergistically enhances nanosonosensitizer-enabled sonodynamic cancer nanotherapeutics
Journal of Nanobiotechnology (2021)