Abstract
Background
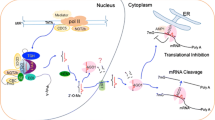
MicroRNAs (miRNAs) have emerged as key regulators of post-transcriptional degradation and/or translational repression in both plants and animals. Increasing evidence has pointed to the important role of intergenic miRNAs in response to environmental stresses; however, detailed information about intronic miRNAs in plants is not clear.
Results
Here, we performed quantitative real-time PCR (qRT-PCR) analysis using transgenic plants to investigate the relationship between intronic miRNAs and their respective host genes in Arabidopsis. Here, we found that three Arabidopsis thaliana intronic miRNAs (miR400, miR838, and miR848) were co-transcribed with their host genes in different organs. Intriguingly, both miR400 and its host gene (At1g32583) were up-regulated during cold stress. Unlike intronic miRNAs in animals, the change of expression levels of plant intronic miRNAs in transgenic lines did not affect the transcriptional levels of their host genes. This result indicates that there is no feedback regulation loop between the intronic miRNAs and their host genes in Arabidopsis. In miRNAs-cropping mutants, the mature miRNAs were significantly reduced, while the expression levels of host genes did not change, suggestting the microprocessor also play roles in intronic miRNA processing.
Conclusions
Together, these results provide the evidence that there is an independent relationship between the processing of intronic miRNAs and their host genes in Arabidopsis.






Similar content being viewed by others
References
Asha S, Nisha J, Soniya EV (2012) In silico characterisation and phylogenetic analysis of two evolutionarily conserved miRNAs (miR166 and miR171) from black pepper (Piper nigrum L.). Plant Mol Biol Rep 31:707–718. https://doi.org/10.1007/s11105-012-0532-5
Aukerman MJ, Sakai H (2003) Regulation of flowering time and floral organ identity by a MicroRNA and its APETALA2-like target genes. Plant Cell 15:2730–2741. https://doi.org/10.1105/tpc.016238
Baskerville S, Bartel DP (2005) Microarray profiling of microRNAs reveals frequent coexpression with neighboring miRNAs and host genes. RNA 11:241–247. https://doi.org/10.1261/rna.7240905
Brown JW, Marshall DF, Echeverria M (2008) Intronic noncoding RNAs and splicing. Trends Plant Sci 13:335–342. https://doi.org/10.1016/j.tplants
Buchan JR, Parker R (2007) Molecular biology the two faces of miRNA. Science 318:1877–1878. https://doi.org/10.1126/science.1152623
Çelik Ö, Akdaş EY (2019) Tissue-specific transcriptional regulation of seven heavy metal stress-responsive miRNAs and their putative targets in nickel indicator castor bean (R. communis L.) plants. Ecotoxicol Environ Saf 170:682–690. https://doi.org/10.1016/j.ecoenv.2018.12.006
Conine CC, Moresco JJ, Gu W, Shirayama M, Conte D Jr, Yates JR 3rd, Mello CC (2013) Argonautes promote male fertility and provide a paternal memory of germline gene expression in C. elegans. Cell 155:1532–1544. https://doi.org/10.1016/j.cell.2013.11.032
Conine CC, Sun F, Song L, Rivera-Pérez JA, Rando OJ (2018) Small RNAs gained during epididymal transit of sperm are essential for embryonic development in mice. Dev Cell 46:470-480.e473. https://doi.org/10.1016/j.devcel.2018.06.024
Djuranovic S, Nahvi A, Green R (2012) miRNA-mediated gene silencing by translational repression followed by mRNA deadenylation and decay. Science 336:237–240. https://doi.org/10.1126/science.1215691
Dong Z, Han MH, Fedoroff N (2008) The RNA-binding proteins HYL1 and SE promote accurate in vitro processing of pri-miRNA by DCL1. Proc Natl Acad Sci USA 105:9970–9975. https://doi.org/10.1073/pnas.0803356105
Fang X, Qi Y (2016) RNAi in plants: an Argonaute-centered view. Plant Cell 28:272–285. https://doi.org/10.1105/tpc.15.00920
Fang Y, Spector DL (2007) Identification of nuclear dicing bodies containing proteins for microRNA biogenesis in living Arabidopsis plants. Curr Biol 17:818–823. https://doi.org/10.1016/j.cub.2007.04.005
Fujioka Y, Utsumi M, Ohba Y, Watanabe Y (2007) Location of a possible miRNA processing site in SmD3/SmB nuclear bodies in Arabidopsis. Plant Cell Physiol 48:12430
Gao X, Qiao Y, Han D, Zhang Y, Ma N (2012) Enemy or partner: relationship between intronic micrornas and their host genes. IUBMB Life 64:835–840. https://doi.org/10.1002/iub.1079
Godnic I, Zorc M, Jevsinek Skok D, Calin GA, Horvat S, Dovc P, Kovac M, Kunej T (2013) Genome-wide and species-wide in silico screening for intragenic MicroRNAs in human, mouse and chicken. PLoS One 8(6):e65165. https://doi.org/10.1371/journal.pone.0065165
Hajieghrari B, Farrokhi N (2020) Investigation on the conserved microRNA genes in higher plants. Plant Mol Biol Rep. https://doi.org/10.1007/s11105-020-01228-9
Han J, Denli AM, Gage FH (2012) The enemy within: intronic miR-26b represses its host gene, ctdsp2, to regulate neurogenesis. Genes Dev 26:6–10. https://doi.org/10.1101/gad.184416.111
Janas MM et al (2011) Feed-forward microprocessing and splicing activities at a microRNA-containing intron. Plos Genet 7:e1002330. https://doi.org/10.1371/journal.pgen.1002330
Jones-Rhoades MW, Bartel DP, Bartel B (2006) MicroRNAs and their regulatory roles in plants. Annu Rev Plant Biol 57:19–53. https://doi.org/10.1146/annurev.arplant.57.032905.105218
Kang L, Han C, Yang G (2020) miR-378 and its host gene Ppargc1β exhibit independent expression in mouse skeletal muscle. Acta Biochimica et Biophysica Sinica 52(8):883–890. https://doi.org/10.1093/abbs/gmaa061
Li Y, Guo Q, Liu P (2021) Dual roles of the Serine/Arginine-rich splicing factor SR45a in promoting and interacting with nuclear cap-binding complex to modulate the salt stress response in Arabidopsis. New Phytol 230(2):641–655. https://doi.org/10.1111/nph.17175
Liu N, Okamura K, Tyler DM, Phillips MD, Chung WJ, Lai EC (2008) The evolution and functional diversification of animal microRNA genes. Cell Res 18:985–996. https://doi.org/10.1038/cr.2008.278
Liu S et al (2020) AtPLATZ2 negatively regulates salt tolerance in Arabidopsis seedlings by directly suppressing the expression of the CBL4/SOS3 and CBL10/SCaBP8 genes. J Exp Bot. https://doi.org/10.1093/jxb/eraa259
Long R, Li M, Li X, Gao Y, Yang Q (2017) A novel miRNA sponge form efficiently inhibits the activity of miR393 and enhances the salt tolerance and ABA insensitivity in Arabidopsis thaliana. Plant Mol Biol Rep 35:1–7. https://doi.org/10.1007/s11105-017-1033-3
Meyers BC et al (2008) Criteria for annotation of plant microRNAs. Plant Cell 20:3186–3190. https://doi.org/10.1105/tpc.108.064311
Morlando M, Ballarino M, Gromak N, Pagano F, Bozzoni I, Proudfoot NJ (2008) Primary microRNA transcripts are processed co-transcriptionally. Nat Struct Mol Biol 15:902–909. https://doi.org/10.1038/nsmb.1475
Reyes JL, Chua NH (2007) ABA induction of miR159 controls transcript levels of two MYB factors during Arabidopsis seed germination. Plant J 49:592–606. https://doi.org/10.1111/j.1365-313X.2006.02980.x
Rodriguez A, Griffiths-Jones S, Ashurst JL, Bradley A (2004) Identification of mammalian microRNA host genes and transcription units. Genome Res 14:1902–1910. https://doi.org/10.1101/gr.2722704
Rogers K, Chen X (2013) Biogenesis, turnover, and mode of action of plant microRNAs. Plant Cell 25:2383–2399. https://doi.org/10.1105/tpc.113.113159
Rupaimoole R, Slack FJ (2017) MicroRNA therapeutics: towards a new era for the management of cancer and other diseases. Nat Rev Drug Discov 16:203–222. https://doi.org/10.1038/nrd.2016.246
Shomron N, Levy C (2009) MicroRNA-biogenesis and pre-mRNA splicing crosstalk. J Biomed Biotechnol 2009:594678. https://doi.org/10.1155/2009/594678
Tang G, Yan J, Gu Y, Qiao M, Fan R, Mao Y, Tang X (2012) Construction of short tandem target mimic (STTM) to block the functions of plant and animal microRNAs. Methods 58:118–125. https://doi.org/10.1016/j.ymeth.2012.10.006
Tang Y, Liu H, Guo S, Wang B, Li Z, Chong K, Xu Y (2018) OsmiR396d affects gibberellin and brassinosteroid signaling to regulate plant architecture in rice. Plant Physiol 176:946–959. https://doi.org/10.1104/pp.17.00964
Verma SS, Sinha R, Rahman MH, Megha S, Deyholos MK, Kav NNV (2014) miRNA-mediated posttranscriptional regulation of gene expression in ABR17-transgenic Arabidopsis thaliana under salt stress. Plant Mol Biol Rep 32:1203–1218. https://doi.org/10.1007/s11105-014-0716-2
Voinnet O (2009) Origin, biogenesis, and activity of plant microRNAs. Cell 136:669–687. https://doi.org/10.1016/j.cell.2009.01.046
Wang F, Zheng T, Hu Z, Wu G, Lang C, Liu R (2020) Overexpression of miR319a altered oil body morphogenesis and lipid content in Arabidopsis seeds. Plant Mol Biol Rep 1–7. https://doi.org/10.1007/s11105-020-01217-y
Wang JW, Czech B, Weigel D (2009) miR156-regulated SPL transcription factors define an endogenous flowering pathway in Arabidopsis thaliana. Cell 138:738–749. https://doi.org/10.1016/j.cell.2009.06.014
Yan K et al (2012) Stress-induced alternative splicing provides a mechanism for the regulation of microRNA processing in Arabidopsis thaliana. Mol Cell 48:521–531. https://doi.org/10.1016/j.molcel.2012.08.032
Yang X, Ren W, Zhao Q, Zhang P, Wu F, He Y (2014) Homodimerization of HYL1 ensures the correct selection of cleavage sites in primary miRNA. Nucleic Acids Res 42:12224–12236. https://doi.org/10.1093/nar/gku907
Ying SY, Chang DC, Lin SL (2018) The microRNA. Methods Mol Bio 1733:1–25. https://doi.org/10.1007/978-1-4939-7601-0_1
Yu B et al (2005) Methylation as a crucial step in plant microRNA biogenesis. Science 307:932–935. https://doi.org/10.1126/science.1107130
Yu Z et al (2019) CEPR2 phosphorylates and accelerates the degradation of PYR/PYLs in Arabidopsis. J Exp Bot 70:5457–5469. https://doi.org/10.1093/jxb/erz302
Zeidler M, Hüttenhofer A, Kress M et al (2020) Intragenic microRNAs autoregulate their host genes in both direct and indirect ways—a cross-species analysis. Cells 9(1):232
Zhang J, Zhang H, Srivastava AK, Pan Y, Bai J, Fang J, Shi H (2018a) Knockdown of rice microRNA166 confers drought resistance by causing leaf rolling and altering stem xylem development. Plant Physiol 176:2082–2094. https://doi.org/10.1104/pp.17.01432
Zhang Y, Yun Z, Gong L, Qu H, Duan X, Jiang Y, Zhu H (2018b) Comparison of miRNA evolution and function in plants and animals. MicroRNA 7:4–10. https://doi.org/10.2174/2211536607666180126163031
Zhao Y, Wang S, Wu W, Li L, Jiang T, Zheng B (2018) Clearance of maternal barriers by pat
Zhu Y et al (2012) MicroRNA-26a/b and their host genes cooperate to inhibit the G1/S transition by activating the pRb protein. Nucleic Acids Res 40:4615–4625. https://doi.org/10.1093/nar/gkr1278
Acknowledgements
The authors would like to thank TopEdit (www.topeditsci.com) for its linguistic assistance during the preparation of this manuscript.
Funding
This work was supported by the National Natural Science Foundation (Grant No. 31870234, 31970292 and 31972357) in China.
Author information
Authors and Affiliations
Contributions
Y.L, Q.G, and K.Y conceived and designed the experiments; Y.L and Q.G. performed the experiments; K.Y, and Y.L made the figures and wrote the article; M.W provided important suggestions; and K.Y, Y.L, and C.Z. supervised and complemented the writing. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Ying Li and Qianhuan Guo contributed equally to this work.
Key Messages
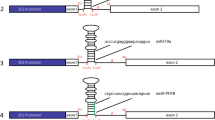
• Arabidopsis intronic miRNAs (miR400, miR838, and miR848) were co-transcribed with their host genes in different tissues.
• miR400 and its host gene were upregulated during cold stress.
• The change of plant intronic miRNAs expression in transgenic lines did not affect the transcriptional levels of their host genes.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, Y., Guo, Q., Wang, M. et al. The Processing and Regulation of Intronic miRNAs Are Independent of Their Host Genes in Arabidopsis. Plant Mol Biol Rep 40, 95–105 (2022). https://doi.org/10.1007/s11105-021-01298-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-021-01298-3